Kinase activity analysis
Last updated: 2023-05-26
Checks: 5 1
Knit directory:
LungCancer_SotilloLab/analysis/
This reproducible R Markdown analysis was created with workflowr (version 1.7.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20221103) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Tracking code development and connecting the code version to the
results is critical for reproducibility. To start using Git, open the
Terminal and type git init in your project directory.
This project is not being versioned with Git. To obtain the full
reproducibility benefits of using workflowr, please see
?wflow_start.
Load packages and dataset
Packages
#package
library(SummarizedExperiment)
library(PHONEMeS) #PHONEMeS-ILP
library(BioNet)
library(OmnipathR)
library(hash)
library(MultiAssayExperiment)
library(PhosR)
library(directPA)
library(tidyverse)
source("../code/utils.R")
knitr::opts_chunk$set(warning = FALSE, message = FALSE, autodep = TRUE)
data(PhosphoSitePlus)
load("../output/allResList_RUN5_timeBased.RData")Using Decoupler for estimating kinase activity (for Figure 6A)
Preprocessing
Construct a mouse version of phonemesPKN
### Construct kinase-substrate interaction network
omnipath_ptm <- get_signed_ptms(enzsub = import_omnipath_enzsub(organism = 10090),
interactions = import_omnipath_interactions(organism = 10090))
omnipath_ptm <- omnipath_ptm[omnipath_ptm$modification %in% c("dephosphorylation", "phosphorylation"), ]
# Filter out ProtMapper
omnipath_ptm_filtered <- omnipath_ptm %>%
dplyr::filter(!(stringr::str_detect(omnipath_ptm$source, "ProtMapper") & n_resources == 1))
# select target (substrate_genesymbol) and source (enzyme_genesymbol)
KSN <- omnipath_ptm_filtered[, c(4, 3)]
# add phosphorylation site to target
KSN$substrate_genesymbol <- paste(KSN$substrate_genesymbol, omnipath_ptm_filtered$residue_type, sep = "_")
KSN$substrate_genesymbol <- paste(KSN$substrate_genesymbol, omnipath_ptm_filtered$residue_offset, sep = "")
# set direction and likelihood of interaction
KSN$mor <- ifelse(omnipath_ptm_filtered$modification == "phosphorylation", 1, -1)
KSN$likelihood <- 1
# we remove ambiguous modes of regulations
KSN$id <- paste(KSN$substrate_genesymbol, KSN$enzyme_genesymbol, sep = "")
KSN <- KSN[!(duplicated(KSN$id) | duplicated(KSN$id, fromLast = TRUE)), ]
KSN <- KSN[, -5]
# rename KSN to fit decoupler format
names(KSN)[1:3] <- c("target", "source", "interaction")
KSN <- KSN[c("source", "interaction", "target")]
phonemesPKN <- KSN %>% filter(interaction ==1)
#rm(KSN, omnipath_ptm, omnipath_ptm_filtered, omnipath_sd, omniR, sif)Using Decoupler to estimate kinase activities
Function for calculating kinase scores
calcKinaseScore <- function(resTab, phonemesPKN, pCut = 0.05, ifFDR = FALSE) {
decoupler_network <- phonemesPKN %>%
dplyr::rename("mor" = interaction) %>%
tibble::add_column("likelihood" = 1)
# get differential phosphorylation sites
resTab <- resTab %>%
arrange(pval) %>% distinct(site, .keep_all = TRUE)
if (ifFDR) {
resTab <- mutate(resTab, pval = adj_pval)
}
inputTab <- filter(resTab, pval <= pCut, site %in% phonemesPKN$target) %>%
select(site, t_statistic) %>% dplyr::rename(t = t_statistic) %>%
data.frame() %>% column_to_rownames("site")
decoupler_network <- decoupleR::intersect_regulons(mat = inputTab,
network = decoupler_network,
.source = source,
.target = target,
minsize = 5)
correlated_regulons <- decoupleR::check_corr(decoupler_network) %>%
dplyr::filter(correlation >= 0.9)
decoupler_network <- decoupler_network %>%
dplyr::filter(!source %in% correlated_regulons$source.2)
kinase_activity <- decoupleR::run_wmean(mat = as.matrix(inputTab),
network = decoupler_network,
sparse = FALSE)
return(kinase_activity)
}Calculate kinase activity score for each time point and comparison
phosRes <- allResList$diffRatio
kinResTab <- lapply(names(phosRes),function(eachTime) {
lapply(unique(phosRes[[eachTime]]$compare), function(eachCompare) {
resTab <- phosRes[[eachTime]] %>% filter(compare == eachCompare)
calcKinaseScore(resTab,phonemesPKN, pCut = 1, ifFDR = FALSE) %>% mutate(time = eachTime, compare = eachCompare)
}) %>% bind_rows()
}) %>% bind_rows() %>%
filter(statistic == "wmean") %>%
select(-statistic, -condition) %>%
mutate(timeCompare = paste0(time, "_", compare))
scoreTab <- select(kinResTab, source, score, timeCompare)
pTab <- select(kinResTab, source, p_value, timeCompare)
#add zero to not estimated values
fullTab <- scoreTab %>%
pivot_wider(names_from = timeCompare, values_from = score) %>%
mutate(across(starts_with("time_"), replace_na, 0)) %>%
pivot_longer(starts_with("time_"), names_to = "timeCompare", values_to = "score" ) %>%
left_join(distinct(kinResTab, timeCompare, time, compare), by = "timeCompare") %>%
left_join(pTab, by = c("source","timeCompare")) %>%
mutate(p_value = ifelse(is.na(p_value),1,p_value)) %>%
dplyr::rename(kinase = "source")Table output of kinase activity scores
writexl::write_xlsx(select(fullTab, kinase, score, time, compare, p_value), "../docs/kinase_activity_decoupler.xlsx")Heatmap plot for kinase activity
plotTab <- mutate(fullTab, sig = ifelse(p_value <=0.05, "*", ""))%>%
mutate(time = paste0(str_remove(time,"time_"), "h"))
ggplot(plotTab, aes(x=time, y = kinase,fill = score)) +
geom_tile() +
geom_text(aes(label = sig), vjust = 0.5) +
facet_wrap(~compare) +
scale_fill_gradient2(low = "blue", high = "red", mid = "white", midpoint = 0) +
scale_x_discrete(expand = c(0,0)) +scale_y_discrete(expand = c(0,0)) +
theme_bw() +
ylab("Kinase")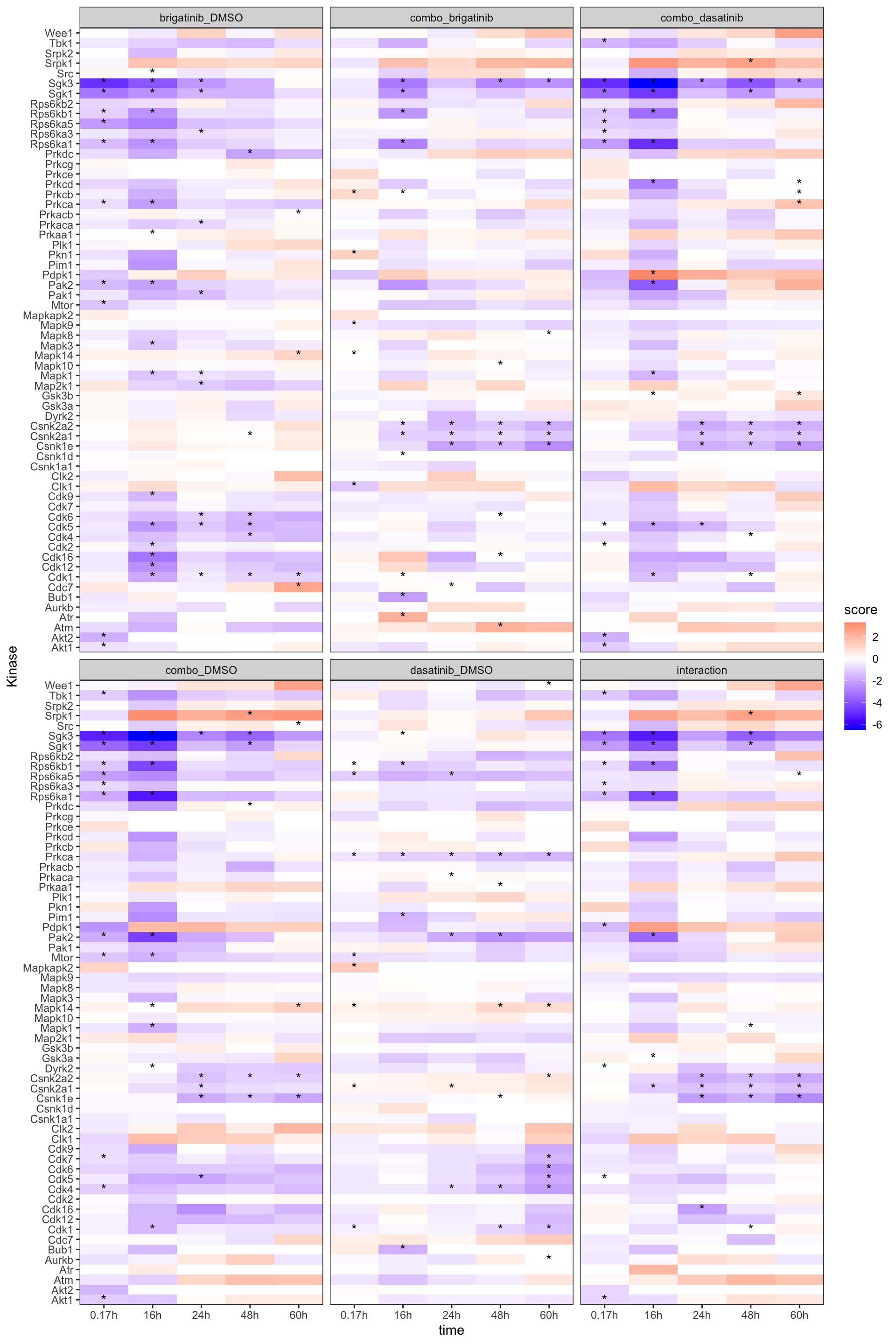
ggsave("../docs/kinase_decoupler_heatmap.pdf", height = 15, width = 10)PDF file: kinase_decoupler_heatmap.pdf
KinasePA analysis for estimating difference in kinase activity scores (perhaps for a supplementary plot)
Phospho at 10min
10 mins
Differential results
resList <- allResList$diffRatio$time_0.17 %>%
filter(compare %in% c("combo_DMSO","brigatinib_DMSO", "dasatinib_DMSO"))
phosTab <- resList %>%
mutate(site = paste0(str_replace(toupper(site),"_",";"),";")) %>%
select(site, t_statistic, compare) %>%
arrange(abs(t_statistic)) %>%
distinct(site, compare,.keep_all = TRUE) %>%
pivot_wider(names_from = compare, values_from = t_statistic) %>%
data.frame() %>% column_to_rownames("site")Pair-wise comparison
Combo versus brigatinib
pdf("../docs/DPA_combo_briga_10min.pdf", height = 9, width = 9)
z1 <- perturbPlot2d(Tc=phosTab[,c(1,2)], annotation=PhosphoSite.mouse, cex=0.5, xlim=c(-8, 8), ylim=c(-8, 8), main="Combo versus brigatinib (10 min)")
dev.off()quartz_off_screen
2 PDF file: DPA_combo_briga_10min.pdf
Combo versus dasatinib
pdf("../docs/DPA_combo_dasa_10min.pdf", height = 9, width = 9)
z1 <- perturbPlot2d(Tc=phosTab[,c(1,3)], annotation=PhosphoSite.mouse, cex=0.5, xlim=c(-8, 8), ylim=c(-8, 8), main="Combo versus dasatinib (10 min)")
dev.off()quartz_off_screen
2 PDF file: DPA_combo_dasa_10min.pdf
16 h
Differential results
resList <- allResList$diffRatio$time_16 %>%
filter(compare %in% c("combo_DMSO","brigatinib_DMSO","dasatinib_DMSO"))
phosTab <- resList %>%
mutate(site = paste0(str_replace(toupper(site),"_",";"),";")) %>%
select(site, t_statistic, compare) %>%
arrange(abs(t_statistic)) %>%
distinct(site, compare,.keep_all = TRUE) %>%
pivot_wider(names_from = compare, values_from = t_statistic) %>%
data.frame() %>% column_to_rownames("site")
phosTab <- phosTab[,c("combo_DMSO","brigatinib_DMSO","dasatinib_DMSO")]Combo versus brigatinib
pdf("../docs/DPA_combo_briga_16h.pdf", height = 9, width = 9)
z1 <- perturbPlot2d(Tc=phosTab[,c(1,2)], annotation=PhosphoSite.mouse, cex=0.5, xlim=c(-8, 8), ylim=c(-8, 8), main="Combo versus brigatinib (16h)")
dev.off()quartz_off_screen
2 PDF file: DPA_combo_briga_16h.pdf
Combo versus dasatinib
pdf("../docs/DPA_combo_dasa_16h.pdf", height = 9, width = 9)
z1 <- perturbPlot2d(Tc=phosTab[,c(1,3)], annotation=PhosphoSite.mouse, cex=0.5, xlim=c(-8, 8), ylim=c(-8, 8), main="Combo versus dasatinib (16h)")
dev.off()quartz_off_screen
2 PDF file: DPA_combo_dasa_16h.pdf
sessionInfo()R version 4.2.0 (2022-04-22)
Platform: x86_64-apple-darwin17.0 (64-bit)
Running under: macOS Big Sur/Monterey 10.16
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/4.2/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.2/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] stats4 stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] forcats_0.5.1 stringr_1.4.1
[3] dplyr_1.0.9 purrr_0.3.4
[5] readr_2.1.2 tidyr_1.2.0
[7] tibble_3.1.8 ggplot2_3.4.1
[9] tidyverse_1.3.2 directPA_1.5
[11] PhosR_1.6.0 MultiAssayExperiment_1.22.0
[13] hash_2.2.6.2 OmnipathR_3.4.7
[15] BioNet_1.56.0 RBGL_1.72.0
[17] graph_1.74.0 PHONEMeS_2.0.1
[19] SummarizedExperiment_1.26.1 Biobase_2.56.0
[21] GenomicRanges_1.48.0 GenomeInfoDb_1.32.2
[23] IRanges_2.30.0 S4Vectors_0.34.0
[25] BiocGenerics_0.42.0 MatrixGenerics_1.8.1
[27] matrixStats_0.62.0
loaded via a namespace (and not attached):
[1] readxl_1.4.0 backports_1.4.1 circlize_0.4.15
[4] workflowr_1.7.0 systemfonts_1.0.4 plyr_1.8.7
[7] igraph_1.3.4 digest_0.6.30 htmltools_0.5.4
[10] viridis_0.6.2 fansi_1.0.3 magrittr_2.0.3
[13] checkmate_2.1.0 memoise_2.0.1 googlesheets4_1.0.0
[16] tzdb_0.3.0 limma_3.52.2 Biostrings_2.64.0
[19] modelr_0.1.8 vroom_1.5.7 prettyunits_1.1.1
[22] colorspace_2.0-3 rvest_1.0.2 blob_1.2.3
[25] rappdirs_0.3.3 textshaping_0.3.6 haven_2.5.0
[28] xfun_0.31 crayon_1.5.2 RCurl_1.98-1.7
[31] jsonlite_1.8.3 glue_1.6.2 ruv_0.9.7.1
[34] gtable_0.3.0 gargle_1.2.0 zlibbioc_1.42.0
[37] XVector_0.36.0 DelayedArray_0.22.0 car_3.1-0
[40] shape_1.4.6 decoupleR_2.2.2 abind_1.4-5
[43] scales_1.2.0 pheatmap_1.0.12 DBI_1.1.3
[46] GGally_2.1.2 rstatix_0.7.0 Rcpp_1.0.9
[49] viridisLite_0.4.0 progress_1.2.2 bit_4.0.4
[52] proxy_0.4-27 preprocessCore_1.58.0 httr_1.4.3
[55] RColorBrewer_1.1-3 calibrate_1.7.7 ellipsis_0.3.2
[58] farver_2.1.1 pkgconfig_2.0.3 reshape_0.8.9
[61] sass_0.4.2 dbplyr_2.2.1 utf8_1.2.2
[64] labeling_0.4.2 tidyselect_1.1.2 rlang_1.0.6
[67] reshape2_1.4.4 later_1.3.0 AnnotationDbi_1.58.0
[70] munsell_0.5.0 cellranger_1.1.0 tools_4.2.0
[73] cachem_1.0.6 cli_3.4.1 generics_0.1.3
[76] RSQLite_2.2.15 statnet.common_4.6.0 broom_1.0.0
[79] evaluate_0.15 fastmap_1.1.0 ggdendro_0.1.23
[82] ragg_1.2.2 yaml_2.3.5 knitr_1.39
[85] bit64_4.0.5 fs_1.5.2 KEGGREST_1.36.3
[88] dendextend_1.16.0 xml2_1.3.3 compiler_4.2.0
[91] rstudioapi_0.13 curl_4.3.2 png_0.1-7
[94] e1071_1.7-11 ggsignif_0.6.3 reprex_2.0.1
[97] bslib_0.4.1 stringi_1.7.8 highr_0.9
[100] logger_0.2.2 lattice_0.20-45 Matrix_1.5-4
[103] vctrs_0.5.2 pillar_1.8.0 lifecycle_1.0.3
[106] jquerylib_0.1.4 GlobalOptions_0.1.2 bitops_1.0-7
[109] httpuv_1.6.6 R6_2.5.1 pcaMethods_1.88.0
[112] promises_1.2.0.1 network_1.17.2 gridExtra_2.3
[115] writexl_1.4.0 MASS_7.3-58 assertthat_0.2.1
[118] rprojroot_2.0.3 withr_2.5.0 GenomeInfoDbData_1.2.8
[121] hms_1.1.1 grid_4.2.0 coda_0.19-4
[124] class_7.3-20 rmarkdown_2.14 carData_3.0-5
[127] googledrive_2.0.0 git2r_0.30.1 ggpubr_0.4.0
[130] lubridate_1.8.0