Correlate proteomic profiling with methylation cluster
Junyan Lu
2020-03-07
Last updated: 2020-03-13
Checks: 5 2
Knit directory: Proteomics/analysis/
This reproducible R Markdown analysis was created with workflowr (version 1.6.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown file has unstaged changes. To know which version of the R Markdown file created these results, you’ll want to first commit it to the Git repo. If you’re still working on the analysis, you can ignore this warning. When you’re finished, you can run wflow_publish to commit the R Markdown file and build the HTML.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20200227) was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
- unnamed-chunk-6
To ensure reproducibility of the results, delete the cache directory correlateMethylationCluster_cache and re-run the analysis. To have workflowr automatically delete the cache directory prior to building the file, set delete_cache = TRUE when running wflow_build() or wflow_publish().
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility. The version displayed above was the version of the Git repository at the time these results were generated.
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can use wflow_publish or wflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Ignored files:
Ignored: .DS_Store
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: analysis/.DS_Store
Ignored: analysis/.Rhistory
Ignored: analysis/compareProteomicsRNAseq_cache/
Ignored: analysis/correlateCLLPD_cache/
Ignored: analysis/correlateGenomic_cache/
Ignored: analysis/correlateGenomic_noBlock_MCLL_cache/
Ignored: analysis/correlateGenomic_noBlock_UCLL_cache/
Ignored: analysis/correlateGenomic_noBlock_cache/
Ignored: analysis/correlateGenomic_removePC_cache/
Ignored: analysis/correlateMIR_cache/
Ignored: analysis/correlateMethylationCluster_cache/
Ignored: analysis/predictOutcome_cache/
Ignored: data/.DS_Store
Ignored: output/.DS_Store
Untracked files:
Untracked: analysis/correlateGenomic_noBlock.Rmd
Untracked: analysis/correlateGenomic_noBlock_MCLL.Rmd
Untracked: analysis/correlateGenomic_noBlock_UCLL.Rmd
Untracked: code/utils.R
Untracked: data/190909_CLL_prot_abund_med_norm.tsv
Untracked: data/190909_CLL_prot_abund_no_norm.tsv
Untracked: data/20190423_Proteom_submitted_samples_bereinigt.xlsx
Untracked: data/20191025_Proteom_submitted_samples_final.xlsx
Untracked: data/LUMOS/
Untracked: data/LUMOS_peptides/
Untracked: data/LUMOS_protAnnotation.csv
Untracked: data/SampleAnnotation_cleaned.xlsx
Untracked: data/facTab_IC50atLeast3New.RData
Untracked: data/gmts/
Untracked: data/mapEnsemble.txt
Untracked: data/mapSymbol.txt
Untracked: data/pyprophet_export_aligned.csv
Untracked: data/timsTOF_protAnnotation.csv
Untracked: output/LUMOS_processed.RData
Untracked: output/proteomic_LUMOS_20200227.RData
Untracked: output/proteomic_timsTOF_20200227.RData
Untracked: output/timsTOF_processed.RData
Unstaged changes:
Modified: analysis/correlateGenomic.Rmd
Modified: analysis/correlateMethylationCluster.Rmd
Modified: analysis/index.Rmd
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the R Markdown and HTML files. If you’ve configured a remote Git repository (see ?wflow_git_remote), click on the hyperlinks in the table below to view them.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| html | b8e0823 | Junyan Lu | 2020-03-10 | Build site. |
| Rmd | c8cb45c | Junyan Lu | 2020-03-10 | update analysis |
Detect differentially expressed proteins
Process datasets
We are interested in intermeiate group specific changes, i.e. changes that do not follow the gradient of LP-IP-HP
Process proteomics data
protMat <- assays(protCLL)[["count"]] #without imputationGet methylation cluster information
designMat <- data.frame(row.names = colnames(protMat),
Mclust = factor(patMeta[match(colnames(protMat),patMeta$Patient.ID),]$Methylation_Cluster,
levels = c("IP","LP","HP")))
designMat <- designMat[!is.na(designMat$Mclust),,drop=FALSE]
protMat <- protMat[,rownames(designMat)]How many sample have methylation cluster information
nrow(designMat)[1] 44Numbers of samples in each cluster
table(designMat$Mclust)
IP LP HP
8 21 15 Differential expression using proDA
Fit the probailistic dropout model
fit <- proDA(protMat, design = ~ Mclust,
col_data = designMat)Test for differentially expressed proteins
resList <- lapply(c("LP","HP"), function(n) {
contra <- paste0("Mclust",n)
resTab <- test_diff(fit, contra) %>%
dplyr::rename(id = name, logFC = diff, t=t_statistic,
P.Value = pval, adj.P.Val = adj_pval) %>%
mutate(name = rowData(protCLL[id,])$hgnc_symbol) %>%
select(name, id, logFC, t, P.Value, adj.P.Val) %>%
arrange(P.Value) %>% mutate(Gene = n) %>%
as_tibble()
}) %>% bind_rows()Identifying IP group specific changes (25% FDR)
ipChange <- filter(resList, adj.P.Val <= 0.25) %>%
select(name,id, logFC, Gene) %>%
spread(key = Gene, value = logFC) %>%
filter(HP*LP >0)How many cases show IP specific changes at 25% FDR?
nrow(ipChange)[1] 7Plot IP-specific changes
plotTab <- protMat[ipChange$id,] %>%
data.frame() %>% rownames_to_column("id") %>%
gather(key = "patID", value = "expr",-id) %>%
mutate(Mclust = designMat[patID,],
IGHV = protCLL[,patID]$IGHV.status,
name = rowData(protCLL[id,])$hgnc_symbol) %>%
mutate(Mclust = factor(Mclust, c("LP","IP","HP")))
ggplot(plotTab, aes(x=Mclust, y = expr, fill = Mclust)) +
geom_boxplot() +
geom_point(aes(col = IGHV)) + facet_wrap(~name, scale = "free") +
#theme(legend.position = "none") +
xlab("Methylation Cluster")Warning: Removed 4 rows containing non-finite values (stat_boxplot).Warning: Removed 4 rows containing missing values (geom_point).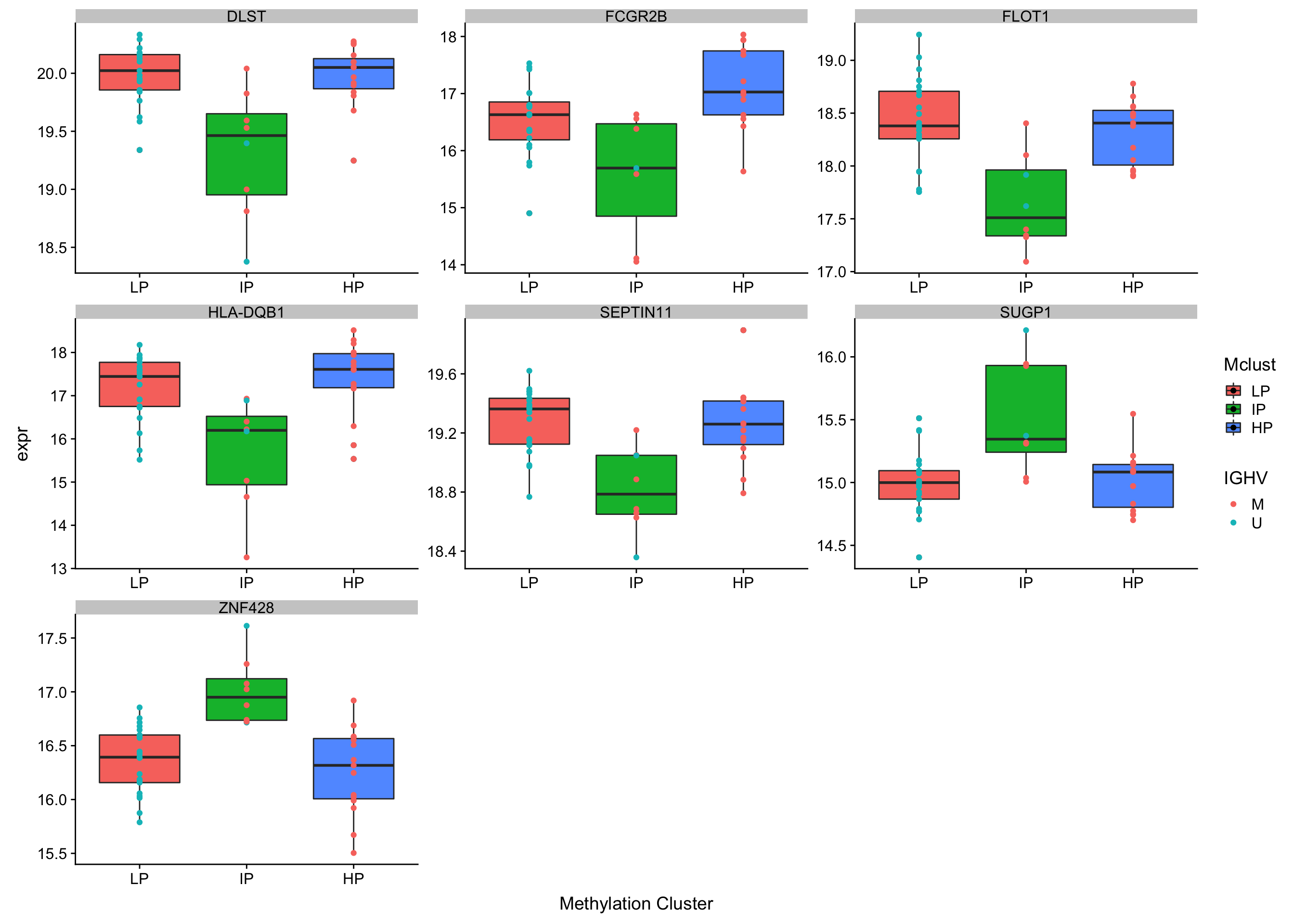
| Version | Author | Date |
|---|---|---|
| b8e0823 | Junyan Lu | 2020-03-10 |
Do the RNA expressions of those 7 genes also correlate with methylation cluster?
dds <- dds[rowSums(counts(dds))>0,]
dds$MClust <- patMeta[match(dds$PatID, patMeta$Patient.ID),]$Methylation_Cluster
ddsSub <- dds[rowData(dds)$symbol %in% ipChange$name, dds$diag %in% "CLL" & !is.na(dds$MClust)]plotTab <- counts(ddsSub, normalized = TRUE) %>% data.frame() %>%
rownames_to_column("id") %>% gather(key = "patID", value = "count",-id) %>%
mutate(MClust = ddsSub[,patID]$MClust,
proteomicSample = ifelse(patID %in% colnames(protCLL),"yes", "no"),
symbol = rowData(ddsSub[id,])$symbol) %>%
mutate(MClust = factor(MClust, levels = c("LP","IP","HP")))
ggplot(plotTab, aes(x=MClust, y = count, fill = MClust)) +
geom_boxplot(outlier.shape = NA) + ggbeeswarm::geom_beeswarm(aes(col = proteomicSample, alpha= proteomicSample)) +
scale_y_log10() + facet_wrap(~symbol,scale = "free", ncol =2) +
scale_color_manual(values = c(yes = "red", no="grey50")) +
scale_alpha_manual(values = c(yes = 1, no = 0.5)) +
xlab("Methylation Cluster")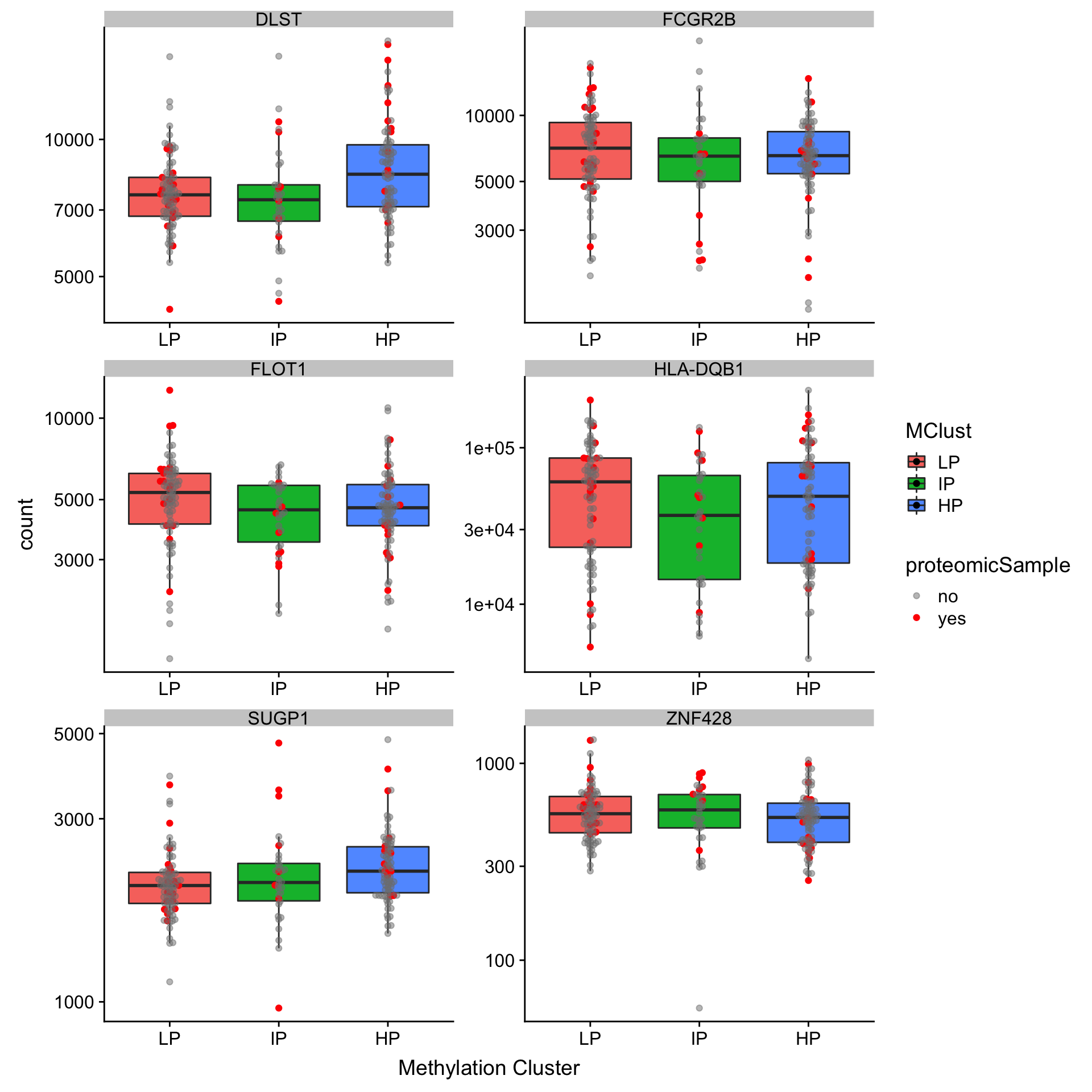
| Version | Author | Date |
|---|---|---|
| b8e0823 | Junyan Lu | 2020-03-10 |
Only six out of seven genes have RNAseq expression and none of them show similar trend as oberserved in proteomic data
Enrichment analysis
Select proteins with IP specific changes (raw p < 0.05)
protList <- filter(resList, P.Value < 0.05) %>%
select(name,id, logFC, Gene) %>%
spread(key = Gene, value = logFC) %>%
filter(HP*LP >0)Rank proteins by the difference to HP and LP group
inputTab <- protList %>% mutate(stat = (HP + LP)/2) %>%
select(name, stat) %>% data.frame() %>% column_to_rownames("name")Enrichment analysis using PAGE
gmts = list(H= "../data/gmts/h.all.v6.2.symbols.gmt",
KEGG= "../data/gmts/c2.cp.kegg.v6.2.symbols.gmt")
enRes <- list()
enRes[["HALLMARK"]] <- runGSEA(inputTab, gmts$H, "page")
enRes[["KEGG"]] <- runGSEA(inputTab, gmts$KEGG, "page")
p <- plotEnrichmentBar(enRes, pCut =0.05, ifFDR= FALSE)Coordinate system already present. Adding new coordinate system, which will replace the existing one.
Coordinate system already present. Adding new coordinate system, which will replace the existing one.plot(p)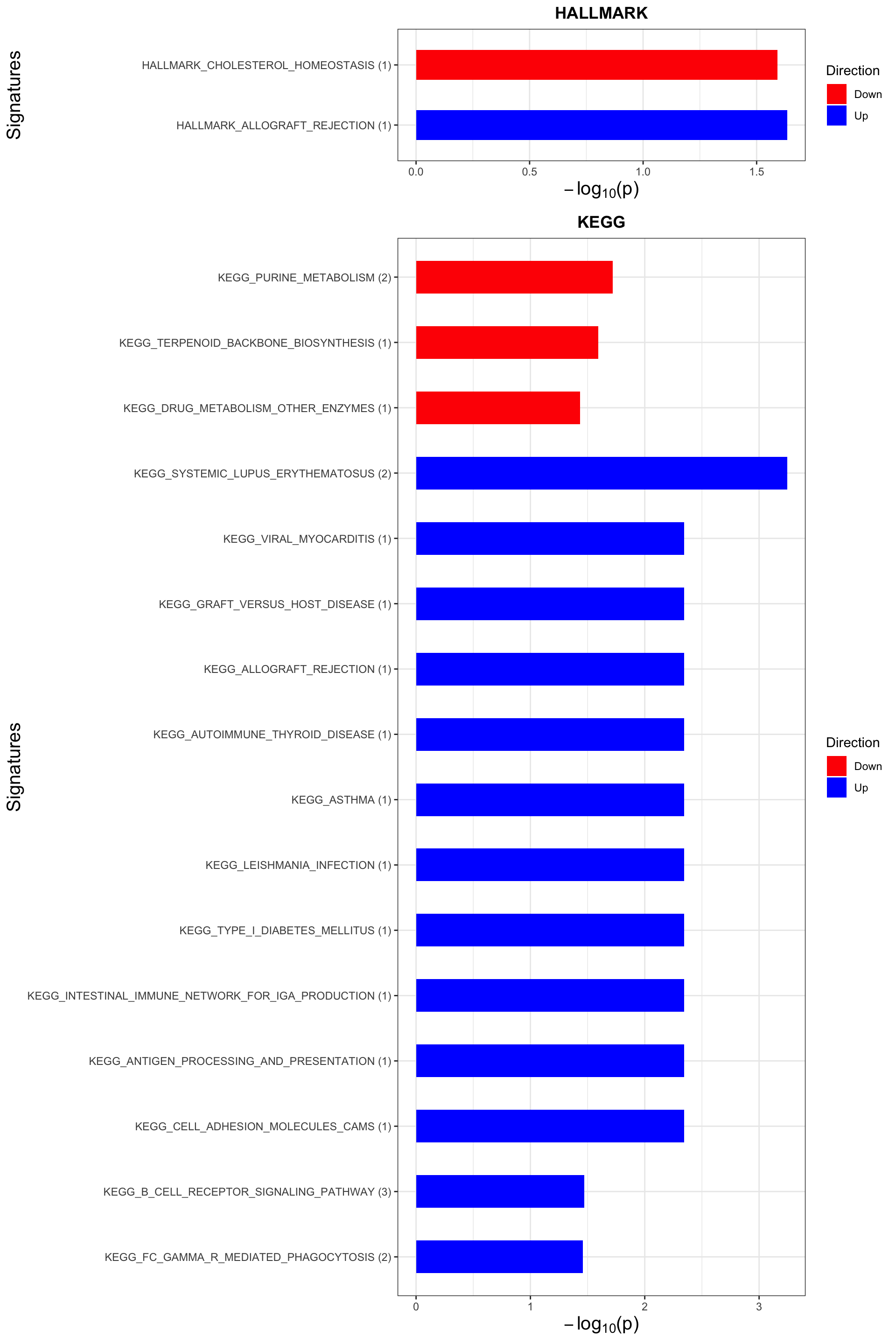
| Version | Author | Date |
|---|---|---|
| b8e0823 | Junyan Lu | 2020-03-10 |
sessionInfo()R version 3.6.0 (2019-04-26)
Platform: x86_64-apple-darwin15.6.0 (64-bit)
Running under: macOS Mojave 10.14.6
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/3.6/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] parallel stats4 stats graphics grDevices utils datasets
[8] methods base
other attached packages:
[1] forcats_0.4.0 stringr_1.4.0
[3] dplyr_0.8.3 purrr_0.3.3
[5] readr_1.3.1 tidyr_1.0.0
[7] tibble_2.1.3 tidyverse_1.3.0
[9] jyluMisc_0.1.5 pheatmap_1.0.12
[11] DESeq2_1.24.0 SummarizedExperiment_1.14.0
[13] DelayedArray_0.10.0 BiocParallel_1.18.0
[15] matrixStats_0.54.0 Biobase_2.44.0
[17] GenomicRanges_1.36.0 GenomeInfoDb_1.20.0
[19] IRanges_2.18.1 S4Vectors_0.22.0
[21] BiocGenerics_0.30.0 piano_2.0.2
[23] proDA_1.1.2 cowplot_0.9.4
[25] ggplot2_3.2.1
loaded via a namespace (and not attached):
[1] shinydashboard_0.7.1 tidyselect_0.2.5 RSQLite_2.1.1
[4] AnnotationDbi_1.46.0 htmlwidgets_1.3 grid_3.6.0
[7] maxstat_0.7-25 munsell_0.5.0 codetools_0.2-16
[10] DT_0.7 withr_2.1.2 colorspace_1.4-1
[13] knitr_1.23 rstudioapi_0.10 ggsignif_0.5.0
[16] labeling_0.3 git2r_0.26.1 slam_0.1-45
[19] GenomeInfoDbData_1.2.1 KMsurv_0.1-5 bit64_0.9-7
[22] rprojroot_1.3-2 vctrs_0.2.0 generics_0.0.2
[25] TH.data_1.0-10 xfun_0.8 sets_1.0-18
[28] R6_2.4.0 ggbeeswarm_0.6.0 locfit_1.5-9.1
[31] bitops_1.0-6 fgsea_1.10.0 assertthat_0.2.1
[34] promises_1.0.1 scales_1.0.0 multcomp_1.4-10
[37] nnet_7.3-12 beeswarm_0.2.3 gtable_0.3.0
[40] sandwich_2.5-1 workflowr_1.6.0 rlang_0.4.1
[43] zeallot_0.1.0 genefilter_1.66.0 cmprsk_2.2-8
[46] splines_3.6.0 lazyeval_0.2.2 acepack_1.4.1
[49] broom_0.5.2 checkmate_1.9.3 yaml_2.2.0
[52] abind_1.4-5 modelr_0.1.5 backports_1.1.4
[55] httpuv_1.5.1 Hmisc_4.2-0 tools_3.6.0
[58] relations_0.6-8 ellipsis_0.2.0 gplots_3.0.1.1
[61] RColorBrewer_1.1-2 Rcpp_1.0.1 base64enc_0.1-3
[64] visNetwork_2.0.7 zlibbioc_1.30.0 RCurl_1.95-4.12
[67] ggpubr_0.2.1 rpart_4.1-15 zoo_1.8-6
[70] haven_2.2.0 cluster_2.1.0 exactRankTests_0.8-30
[73] fs_1.3.1 magrittr_1.5 data.table_1.12.2
[76] openxlsx_4.1.0.1 reprex_0.3.0 survminer_0.4.4
[79] mvtnorm_1.0-11 whisker_0.3-2 hms_0.5.2
[82] shinyjs_1.0 mime_0.7 evaluate_0.14
[85] xtable_1.8-4 XML_3.98-1.20 rio_0.5.16
[88] readxl_1.3.1 gridExtra_2.3 compiler_3.6.0
[91] KernSmooth_2.23-15 crayon_1.3.4 htmltools_0.3.6
[94] later_0.8.0 Formula_1.2-3 geneplotter_1.62.0
[97] lubridate_1.7.4 DBI_1.0.0 dbplyr_1.4.2
[100] MASS_7.3-51.4 Matrix_1.2-17 car_3.0-3
[103] cli_1.1.0 marray_1.62.0 gdata_2.18.0
[106] igraph_1.2.4.1 pkgconfig_2.0.2 km.ci_0.5-2
[109] foreign_0.8-71 xml2_1.2.2 annotate_1.62.0
[112] vipor_0.4.5 XVector_0.24.0 drc_3.0-1
[115] rvest_0.3.5 digest_0.6.19 rmarkdown_1.13
[118] cellranger_1.1.0 fastmatch_1.1-0 survMisc_0.5.5
[121] htmlTable_1.13.1 curl_3.3 shiny_1.3.2
[124] gtools_3.8.1 lifecycle_0.1.0 nlme_3.1-140
[127] jsonlite_1.6 carData_3.0-2 limma_3.40.2
[130] pillar_1.4.2 lattice_0.20-38 httr_1.4.1
[133] plotrix_3.7-6 survival_2.44-1.1 glue_1.3.1
[136] zip_2.0.2 bit_1.1-14 stringi_1.4.3
[139] blob_1.1.1 latticeExtra_0.6-28 caTools_1.17.1.2
[142] memoise_1.1.0