Differential expression analysis on phosphorylation level (RUN5, no normalization by Spectronaut, vst normalization in R)
Last updated: 2023-02-14
Checks: 5 1
Knit directory:
LungCancer_SotilloLab/analysis/
This reproducible R Markdown analysis was created with workflowr (version 1.7.0). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20221103) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Tracking code development and connecting the code version to the
results is critical for reproducibility. To start using Git, open the
Terminal and type git init in your project directory.
This project is not being versioned with Git. To obtain the full
reproducibility benefits of using workflowr, please see
?wflow_start.
Load packages and dataset
Packages
#package
library(SummarizedExperiment)
library(MultiAssayExperiment)
library(proDA)
library(UpSetR)
library(tidyverse)
source("../code/utils.R")
knitr::opts_chunk$set(warning = FALSE, message = FALSE, autodep = TRUE)Pre-processed data
load("../output/processedData_RUN5.RData")
#load saved result list
load("../output/allResList_RUN5_timeBased.RData")
#List of mitochondiral genes
mitoList <- readxl::read_xls("../data/Mouse.MitoCarta3.0.xls", sheet = 2)$Symbol
#geneset files
gmts <- list(Hallmark = "../data/gmts/mh.all.v2022.1.Mm.symbols.gmt",
CanonicalPathway = "../data/gmts/m2.cp.v2022.1.Mm.symbols.gmt",
Kinase = "../data/gmts/Kinase_substrate_noSite.gmt")
#kinase-substrate
kins <- list(Kinase = "../data/gmts/Kinase_substrate.gmt")Differential expression at 10min (0.17 hour) for each drug
Preprocessing
filterList <- list(time = c(0.17,0))
fpeSub <- preprocessPhos(maeData, filterList, missCut = 0.5, transform = "vst", normalize = TRUE)[1] "Number of proteins and samples:"
[1] 5360 24Plot value distribution
plotTab <- jyluMisc::sumToTidy(fpeSub)
ggplot(plotTab, aes(x=sample, y=Intensity)) +
geom_boxplot(aes(fill = drug)) +
theme(axis.text.x = element_text(angle = 90, hjust = 1, vjust = 0.5))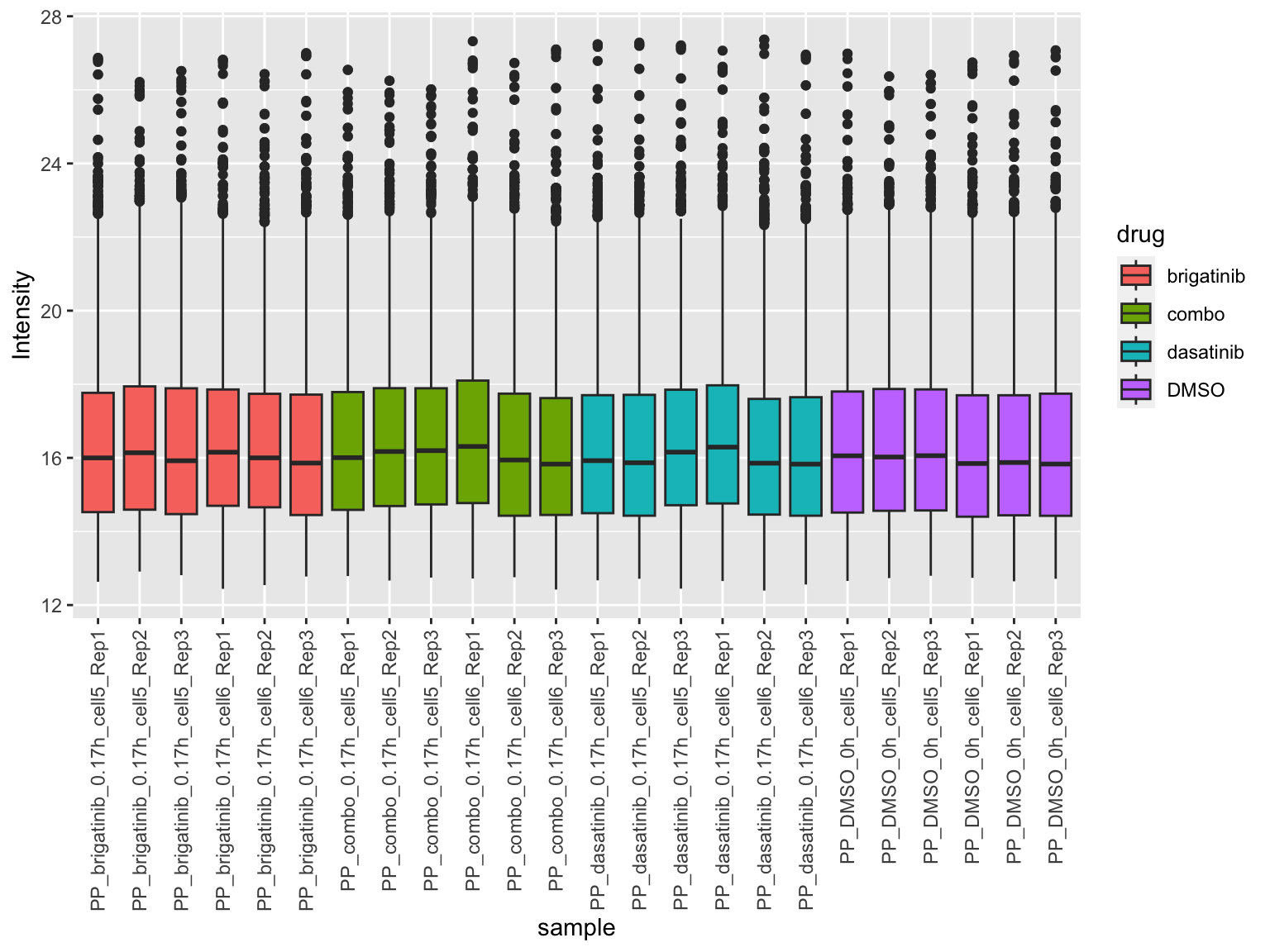 ### PCA
### PCA
PC1 versus PC2
plotPCA(fpeSub, assayName = "imputed", "PC1", "PC2", topVar = 2000, label ="replicate")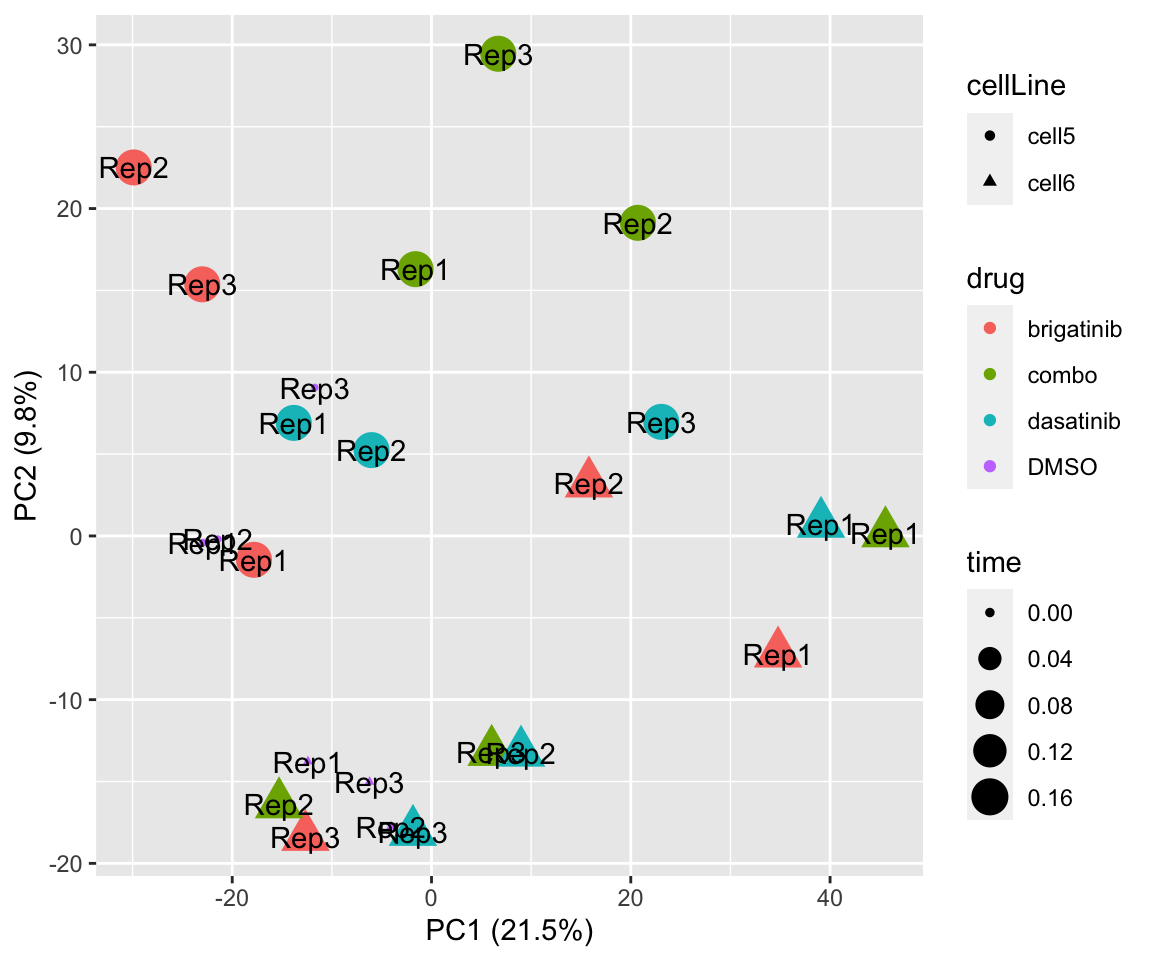 Potential problem with 1 replicate
Potential problem with 1 replicate
PC3 versus PC4
plotPCA(fpeSub, assayName = "imputed", "PC3", "PC4", topVar = 2000, label = "replicate")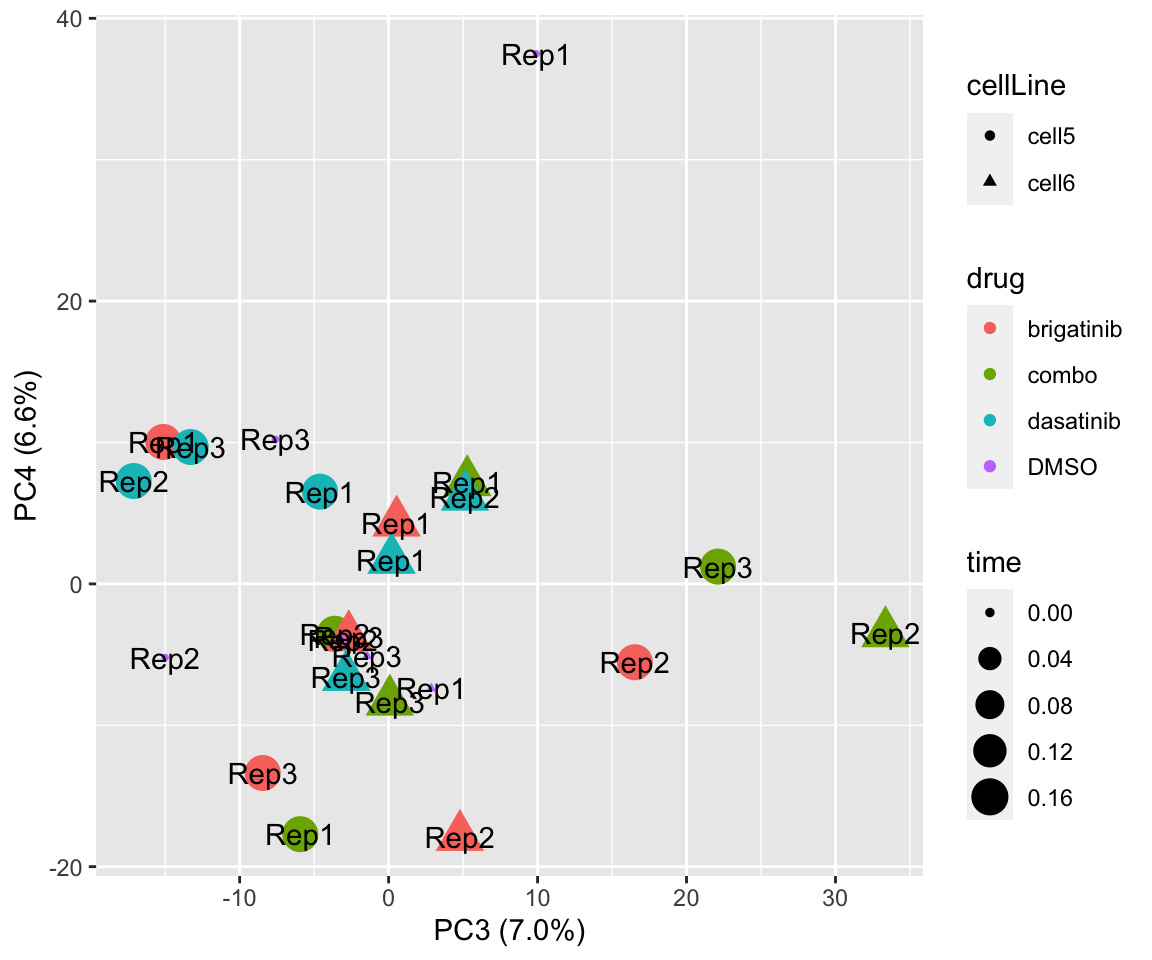
Differential expression using proDA
Use saved results
resTab <- allResList$diffPhos$time_0.17 %>% filter(compare !="interaction")Table of significant associations (10% FDR)
resTab.sig <- filter(resTab, adj_pval <= 0.1)
resTab.sig %>% mutate(across(where(is.numeric), formatC, digits=2)) %>% DT::datatable()P-value histogram for each comparison
ggplot(resTab, aes(x=pval)) +
geom_histogram(bins = 20, fill = "lightblue", color = "grey50") +
facet_wrap(~compare)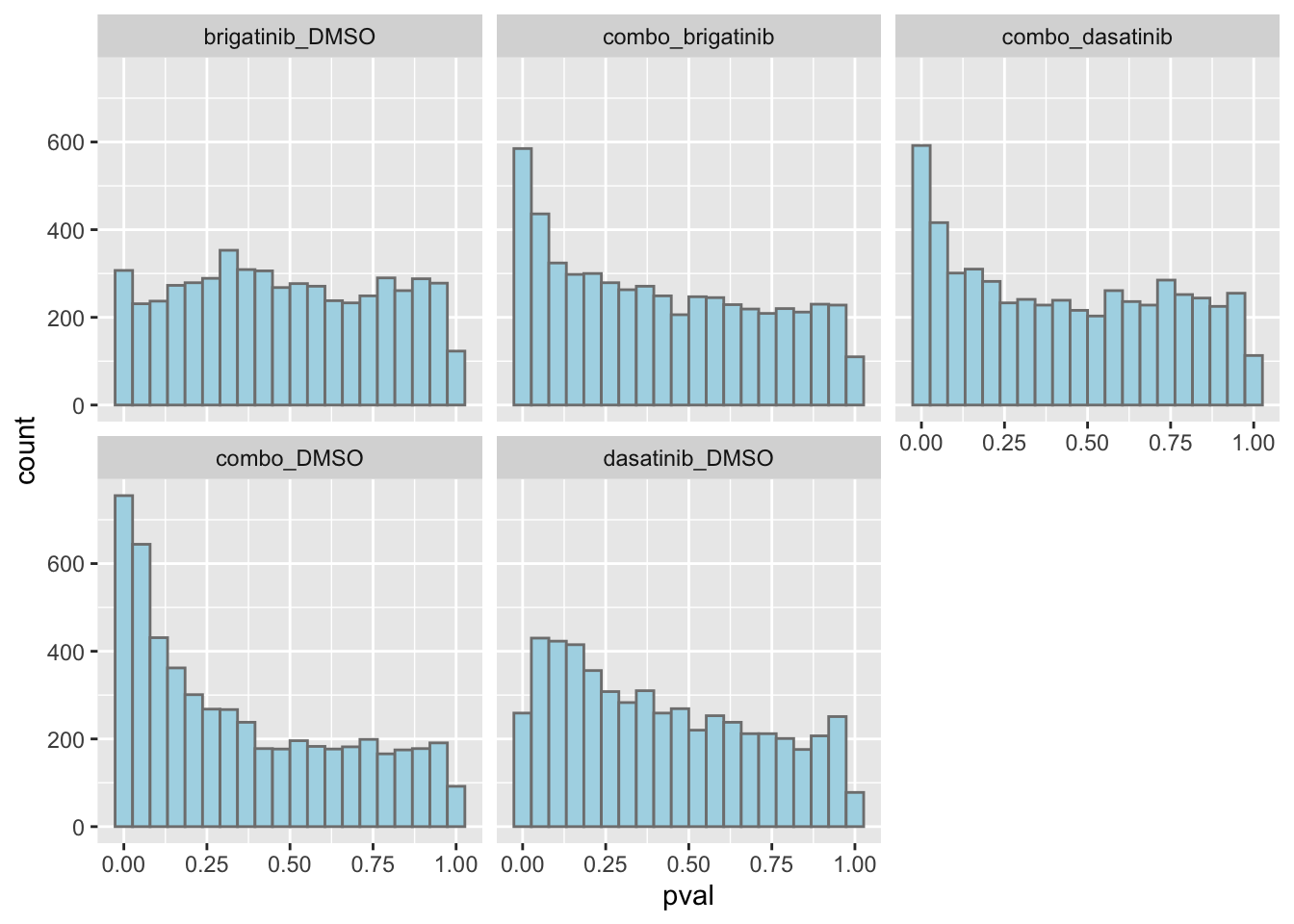
Number of DE proteins for each comparison (10% FDR)
sumTab <- filter(resTab, adj_pval < 0.1) %>%
mutate(direction = ifelse(diff>0, "Up","Down")) %>%
group_by(compare, direction) %>%
summarise(n=length(name))
ggplot(sumTab, aes(x=compare, y=n)) +
geom_bar(aes(fill = direction), stat = "identity", position = "dodge") +
coord_flip() +
geom_text(aes(label =n)) +
facet_wrap(~direction, ncol=2, scales = "free_x") +
xlab("Number")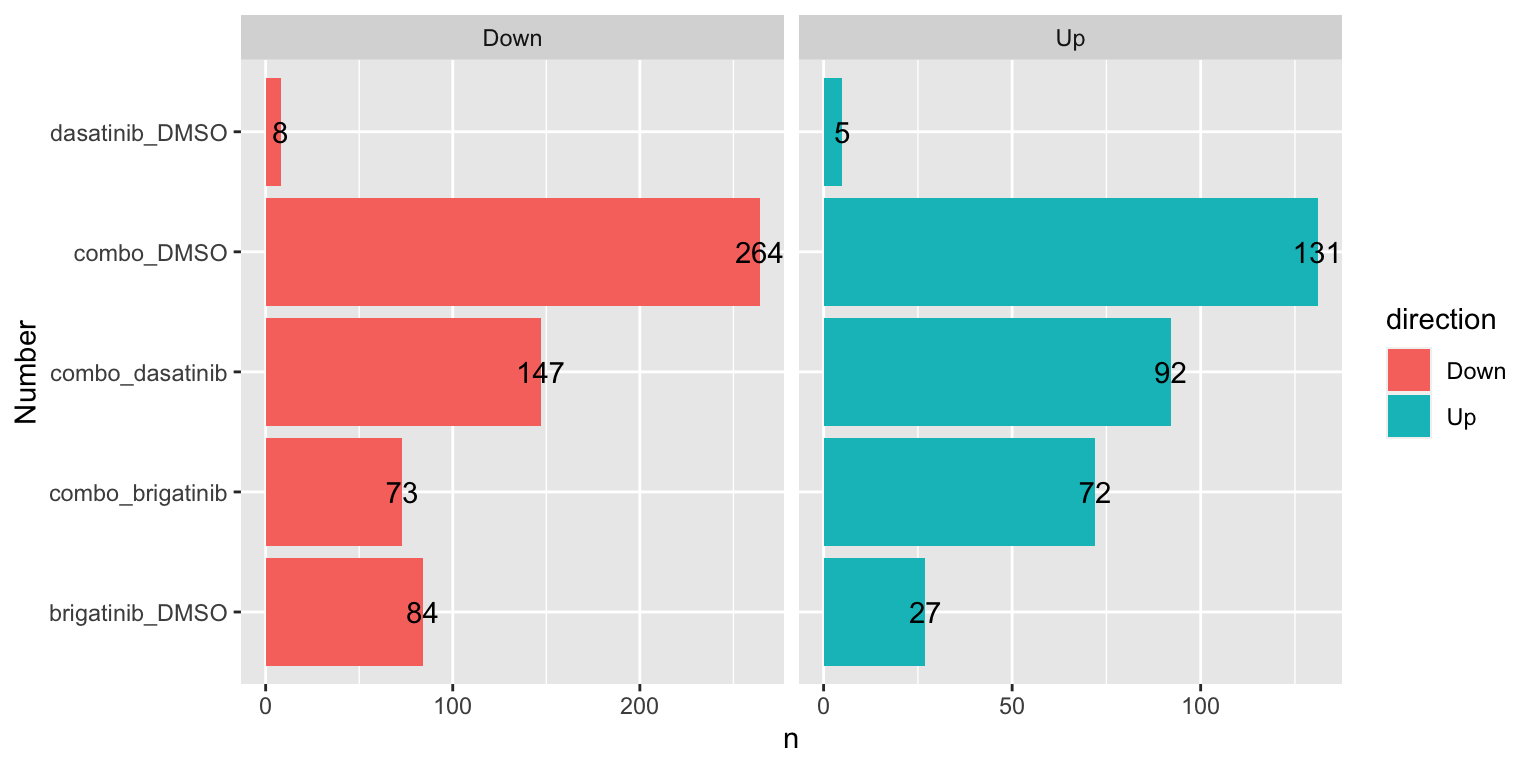
Heatmap of DE proteins for each treatment
Brigatinib versus DMSO
plotProteinHeatmap(resTab, fpeSub, "brigatinib_DMSO", fdrCut = 0.1, ifFDR = TRUE)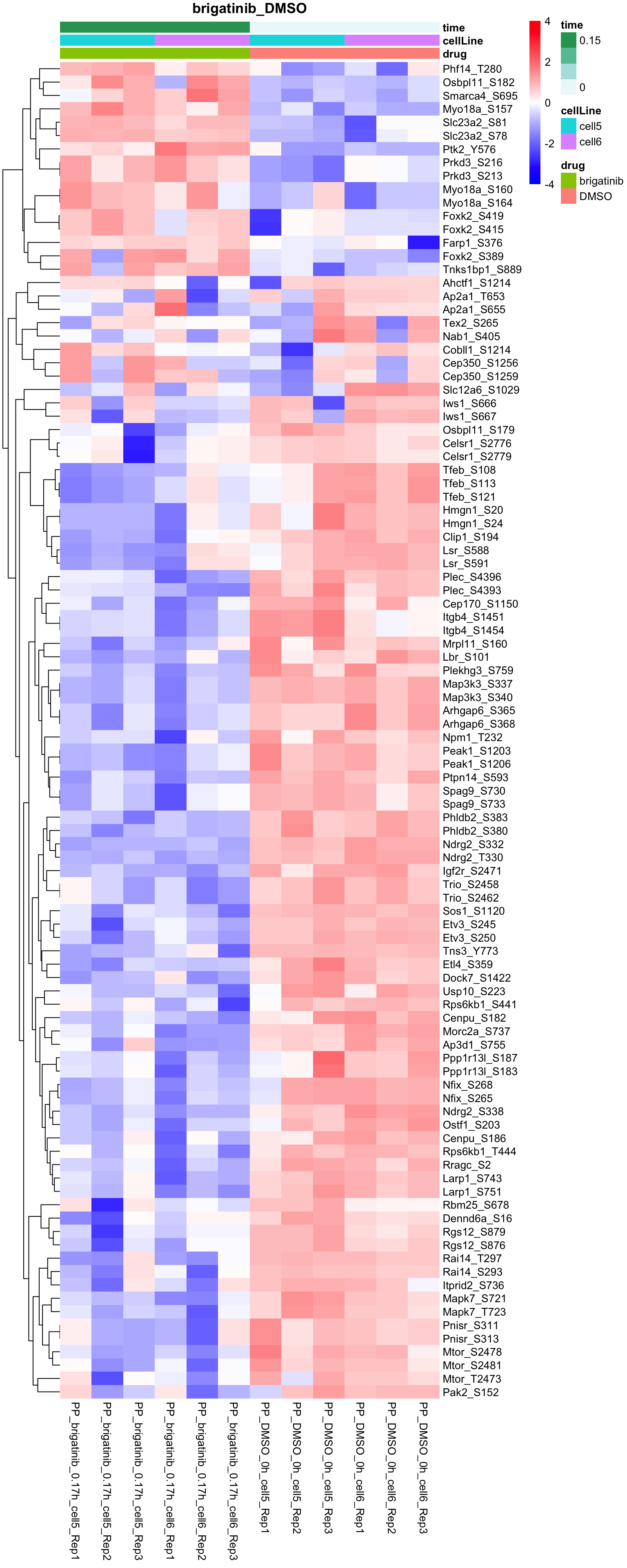
Dasatinib versus DMSO
plotProteinHeatmap(resTab, fpeSub, "dasatinib_DMSO", fdrCut = 0.1, ifFDR =TRUE)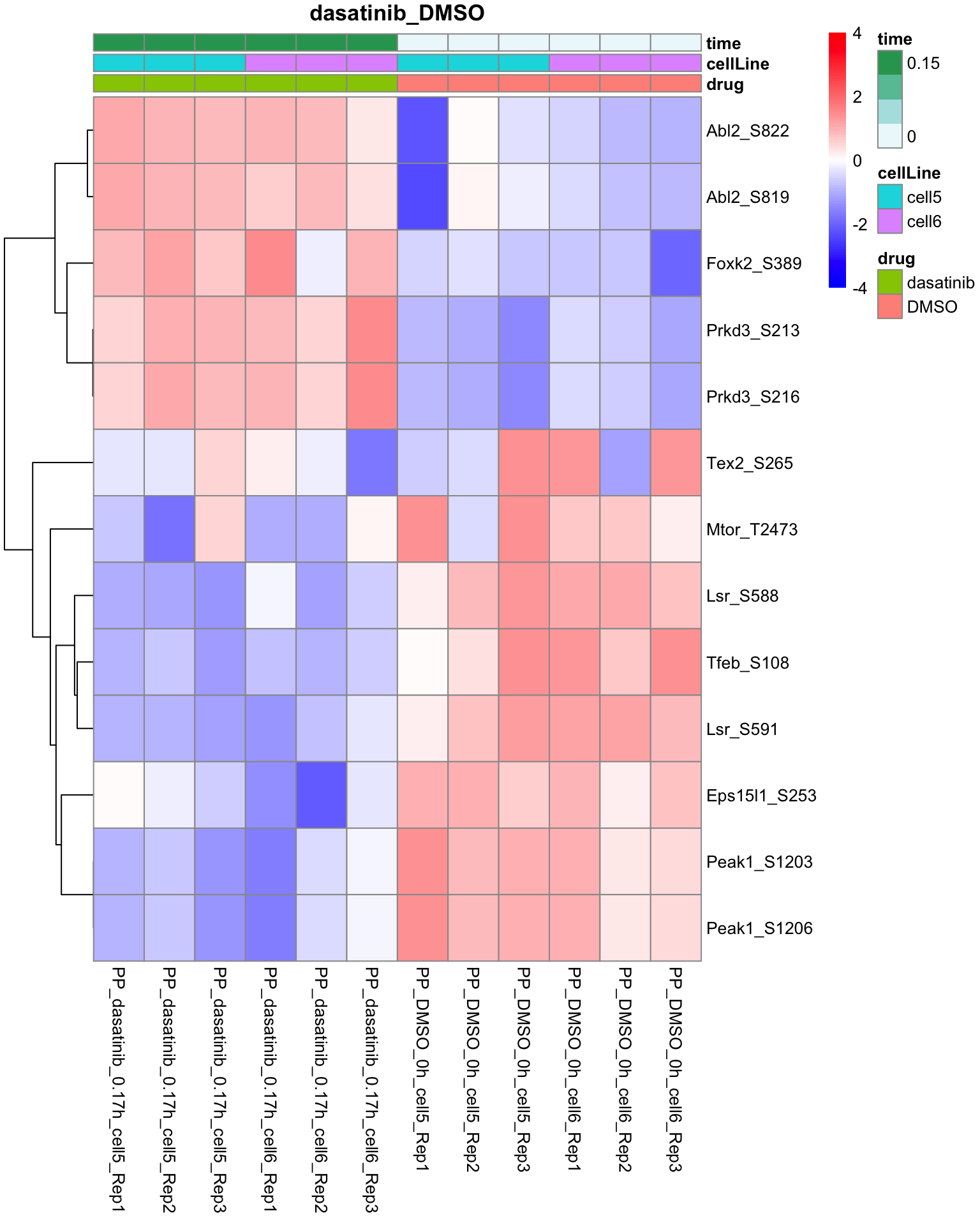
Combo versus DMSO
plotProteinHeatmap(resTab, fpeSub, "combo_DMSO", fdrCut = 0.1, ifFDR = TRUE)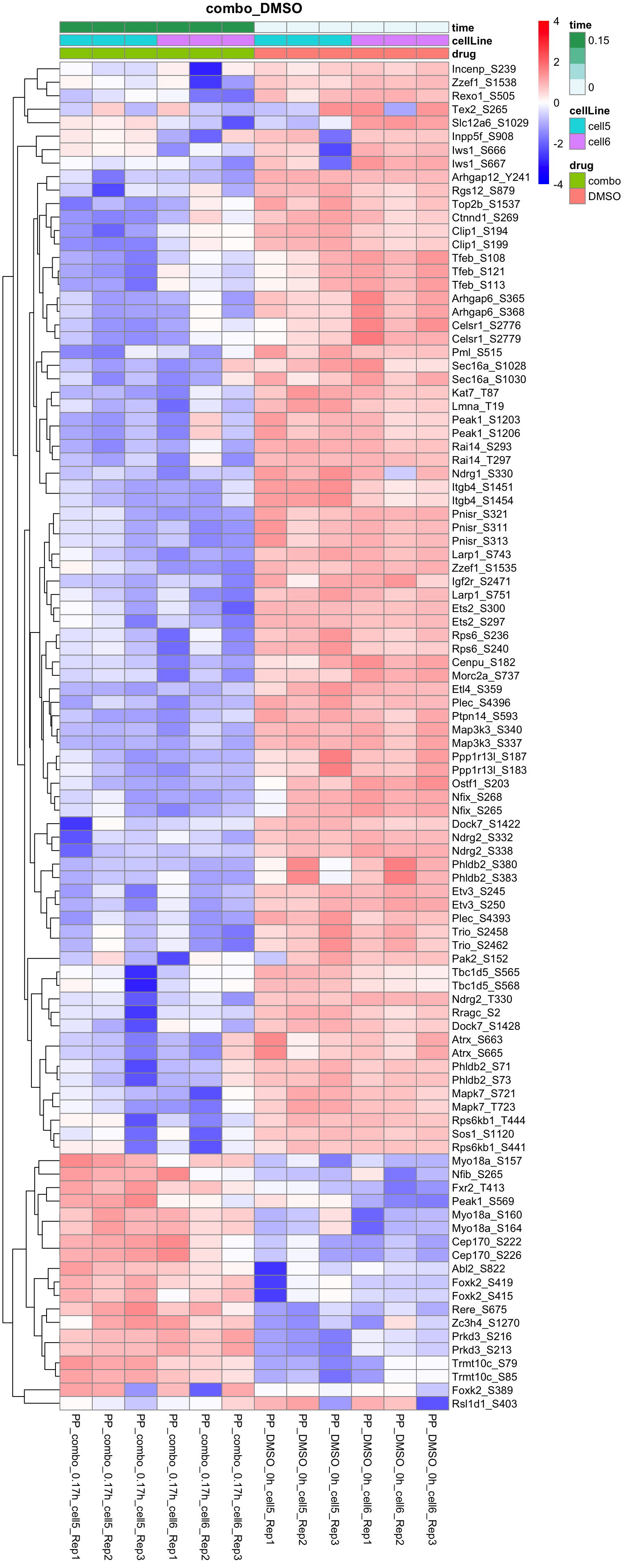
Combo versus brigatinib
plotProteinHeatmap(resTab, fpeSub, "combo_brigatinib", fdrCut = 0.1, ifFDR = TRUE)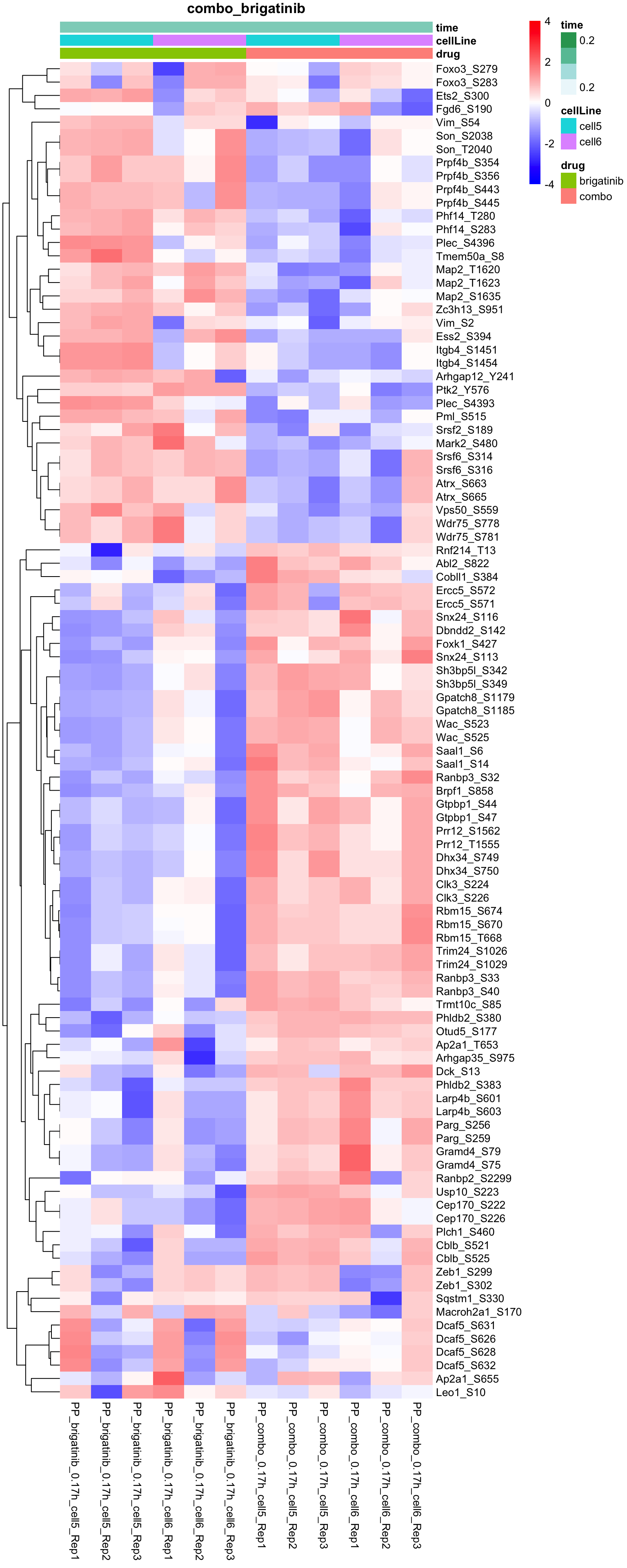
Combo versus dasatinib
plotProteinHeatmap(resTab, fpeSub, "combo_dasatinib", fdrCut = 0.1, ifFDR = TRUE)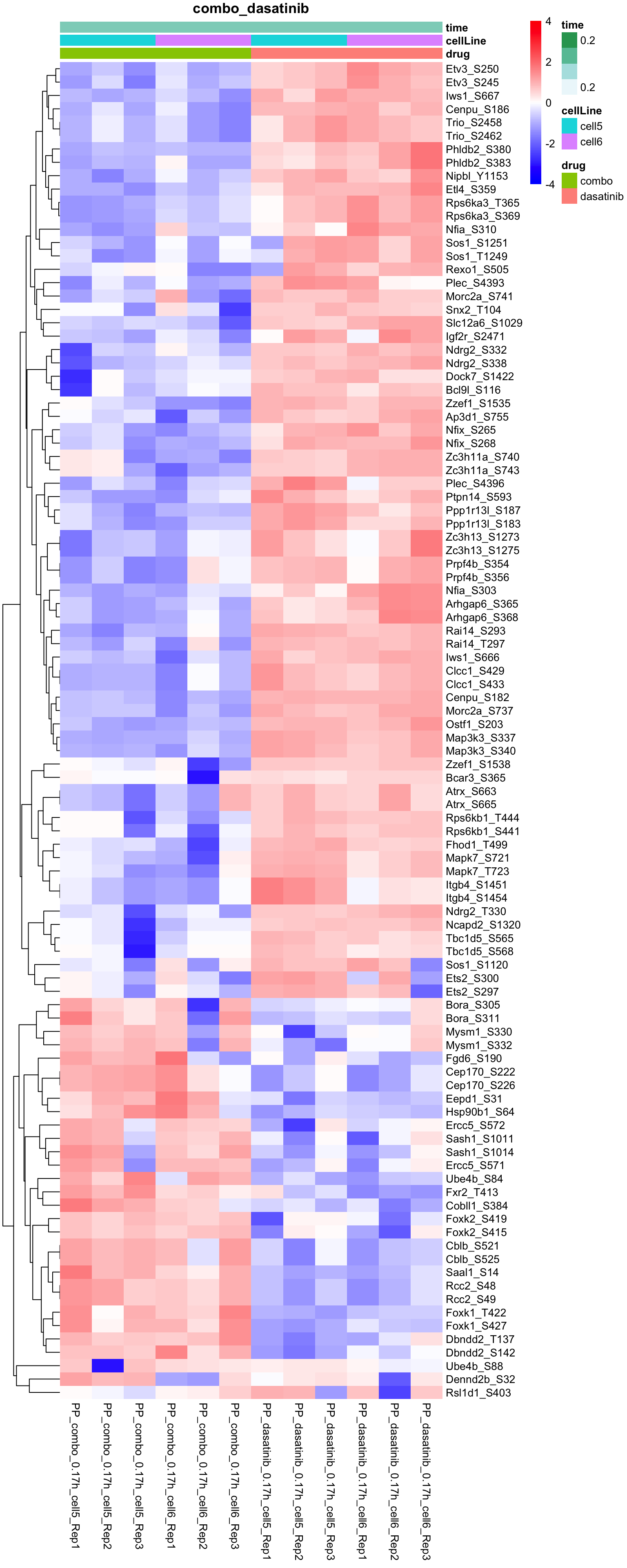
Overlap of DE proteins
Up regulated
deList <- lapply(unique(resTab$compare), function(x) {
filter(resTab, compare == x,adj_pval<0.1, diff >0)$name
})
names(deList) <- unique(resTab$compare)
upset(fromList(deList))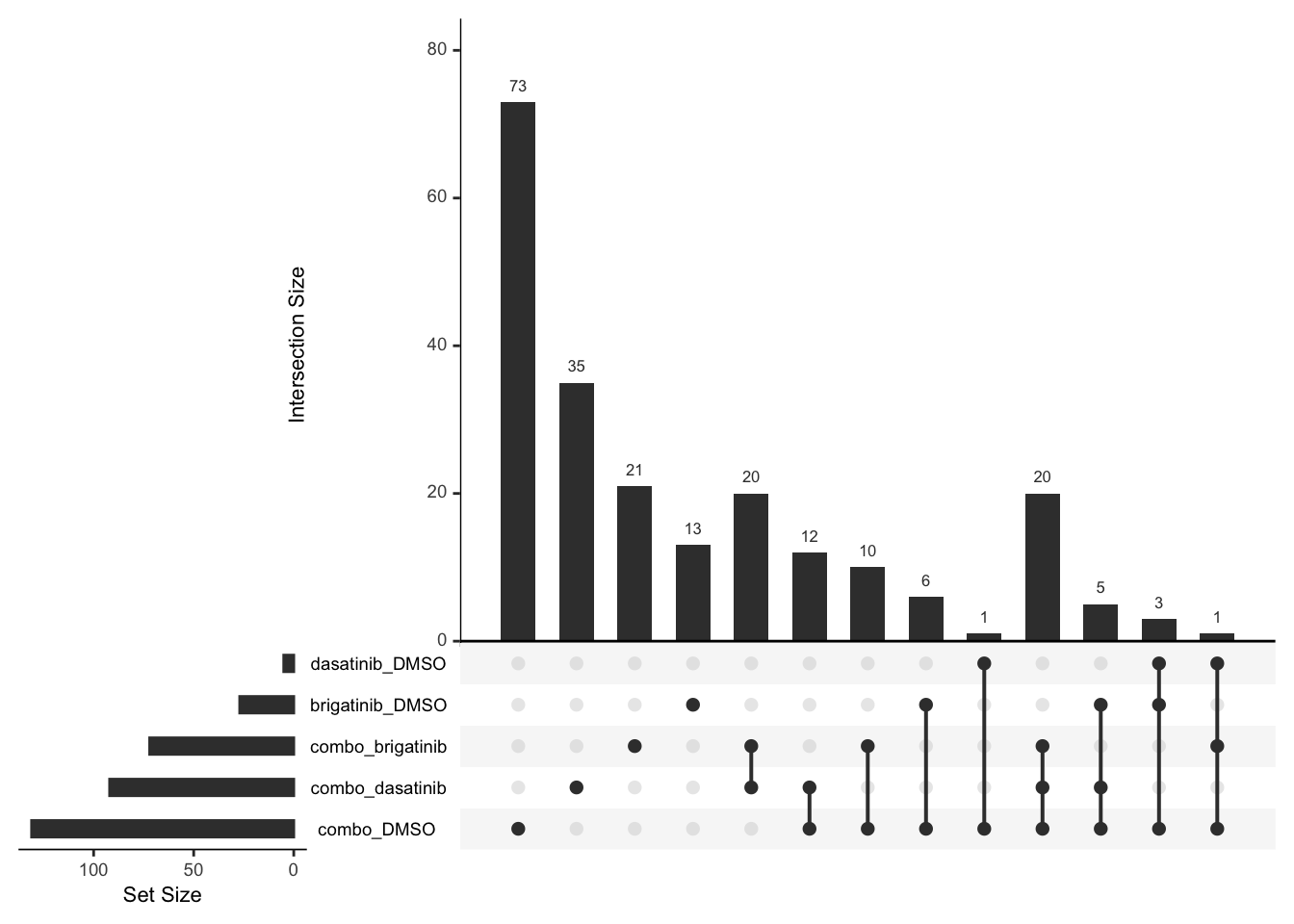
Down regulated
deList <- lapply(unique(resTab$compare), function(x) {
filter(resTab, compare == x, adj_pval <= 0.1, diff <0)$name
})
names(deList) <- unique(resTab$compare)
upset(fromList(deList))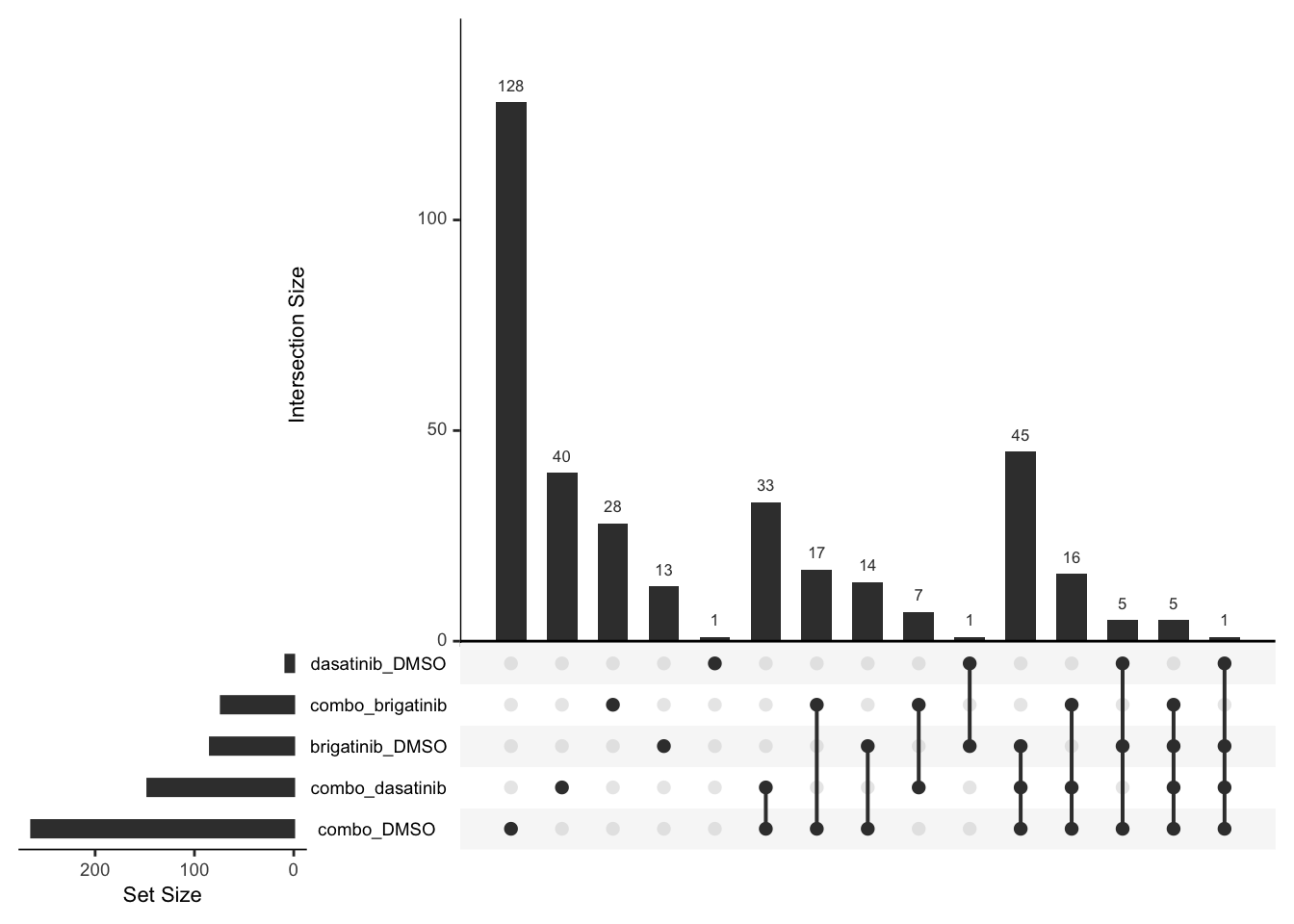
Gene set enrichment analysis
Define useful genesets
plotList <- runGeneSetEnrichment(resTab, gmts, genePCut = 1, pCutSet = 0.25, setFdr = TRUE, method="gsea", collapsePathway = TRUE)[1] "Testing for: Hallmark"
[1] "Condition: dasatinib_DMSO"
[1] "Condition: brigatinib_DMSO"
[1] "Condition: combo_DMSO"
[1] "Condition: combo_brigatinib"
[1] "Condition: combo_dasatinib"
[1] "No sets passed the criteria"
[1] "No sets passed the criteria"
[1] "Testing for: CanonicalPathway"
[1] "Condition: dasatinib_DMSO"
[1] "Condition: brigatinib_DMSO"
[1] "Condition: combo_DMSO"
[1] "Condition: combo_brigatinib"
[1] "Condition: combo_dasatinib"
[1] "No sets passed the criteria"
[1] "Testing for: Kinase"
[1] "Condition: dasatinib_DMSO"
[1] "Condition: brigatinib_DMSO"
[1] "Condition: combo_DMSO"
[1] "Condition: combo_brigatinib"
[1] "Condition: combo_dasatinib"
[1] "No sets passed the criteria"
[1] "No sets passed the criteria"25% FDR is used to explore more pathways
Hallmarks
plotList$Hallmark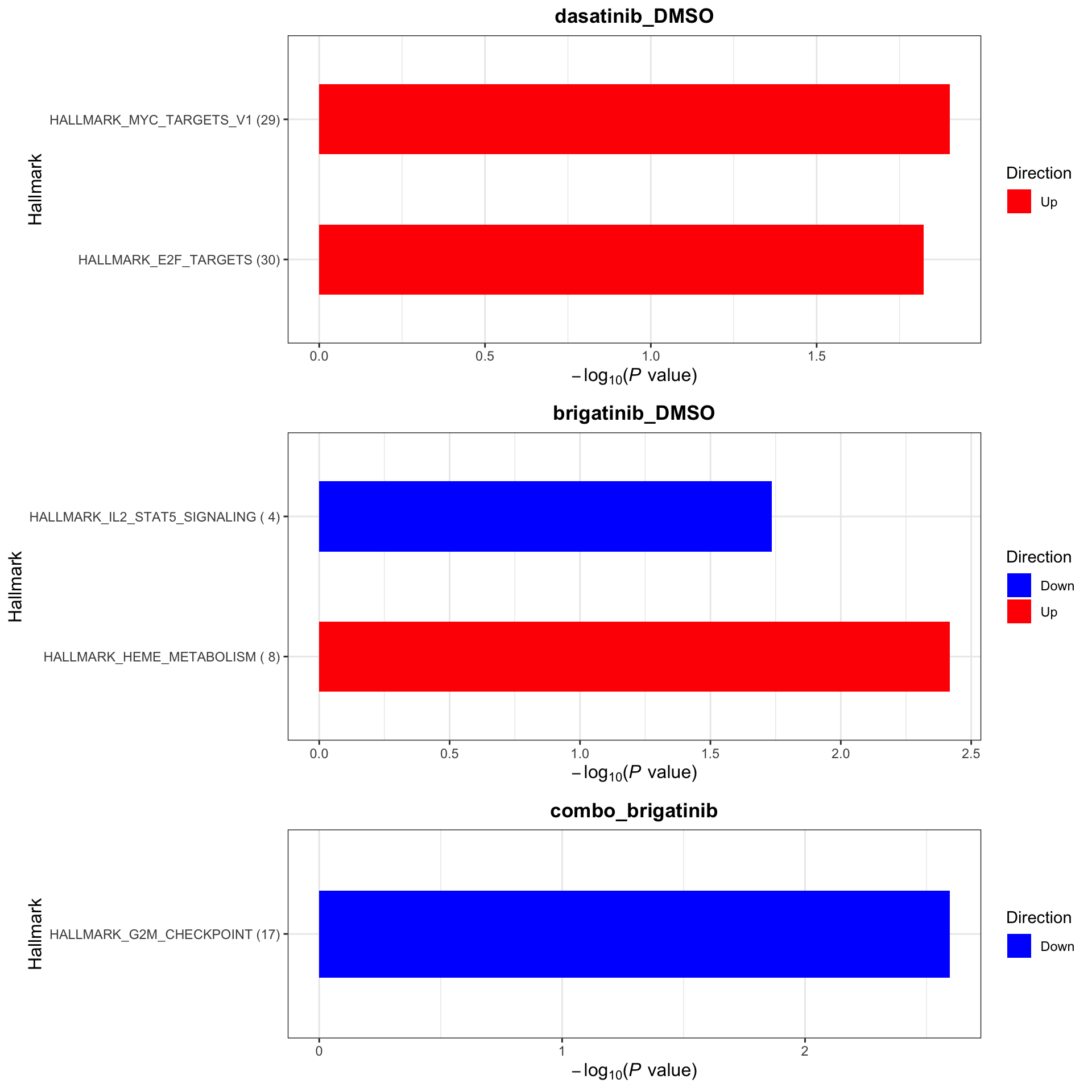
Canonical pathways
plotList$CanonicalPathway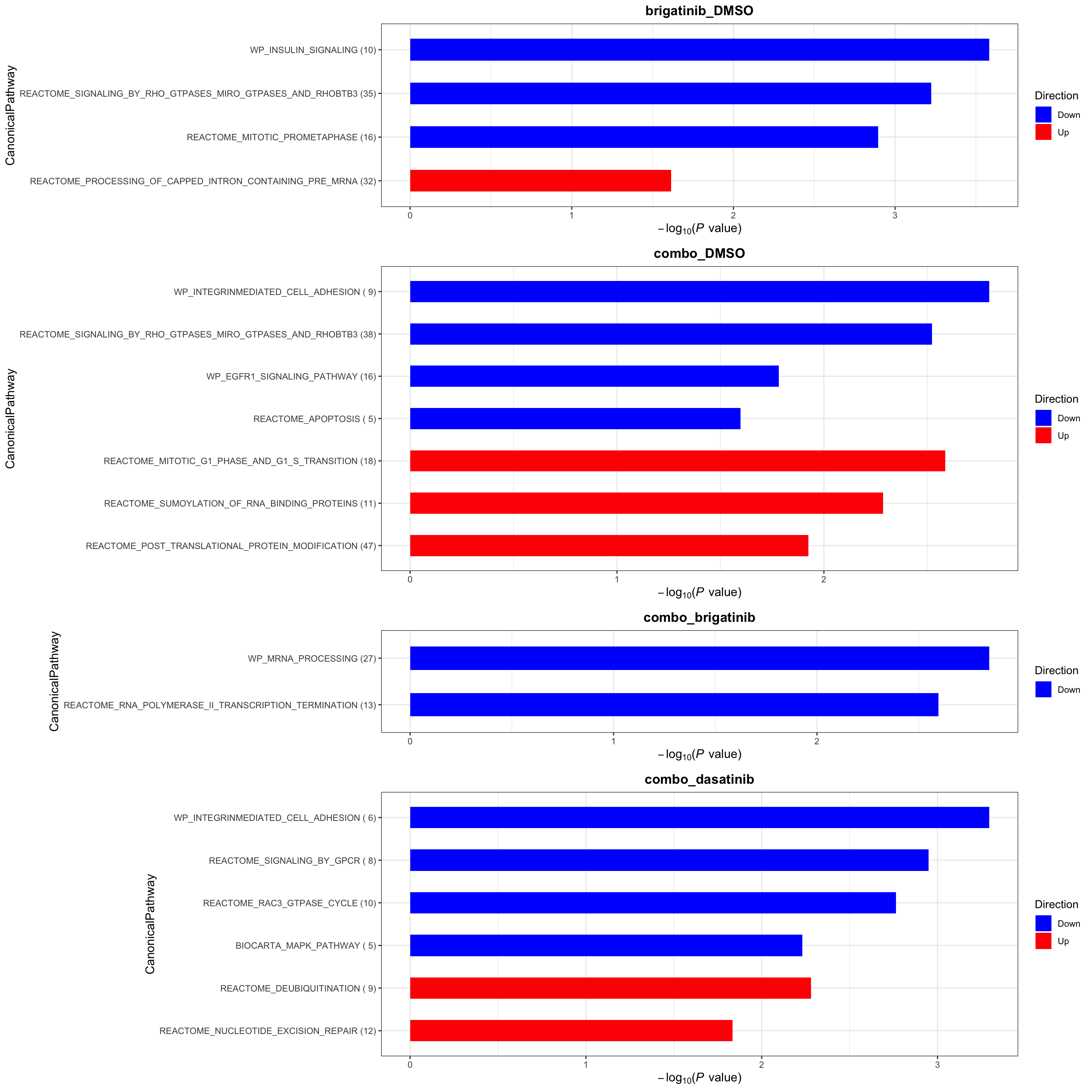
List of leading edge genes for each pathway
Leading edges genes are not necessarily significantly differentially expressed, but they contribute most to the enrichment analysis. Please see explaination of leading edge genes on this page: https://www.gsea-msigdb.org/gsea/doc/GSEAUserGuideFrame.html
DT::datatable(plotList$leadingEdgeGene)writexl::write_xlsx(plotList$leadingEdgeGene, path = "../docs/enrichment_tables/phos_gsea_0.16_all.xlsx")Site-unspecific Kinase enrichment
On protein level, site specificity is not considered. This is because many phosphorylation sites lack the up-stream kinase annotation in the database. We can use a less stringent criteria to define kinase-substrate relation ship
plotList$Kinase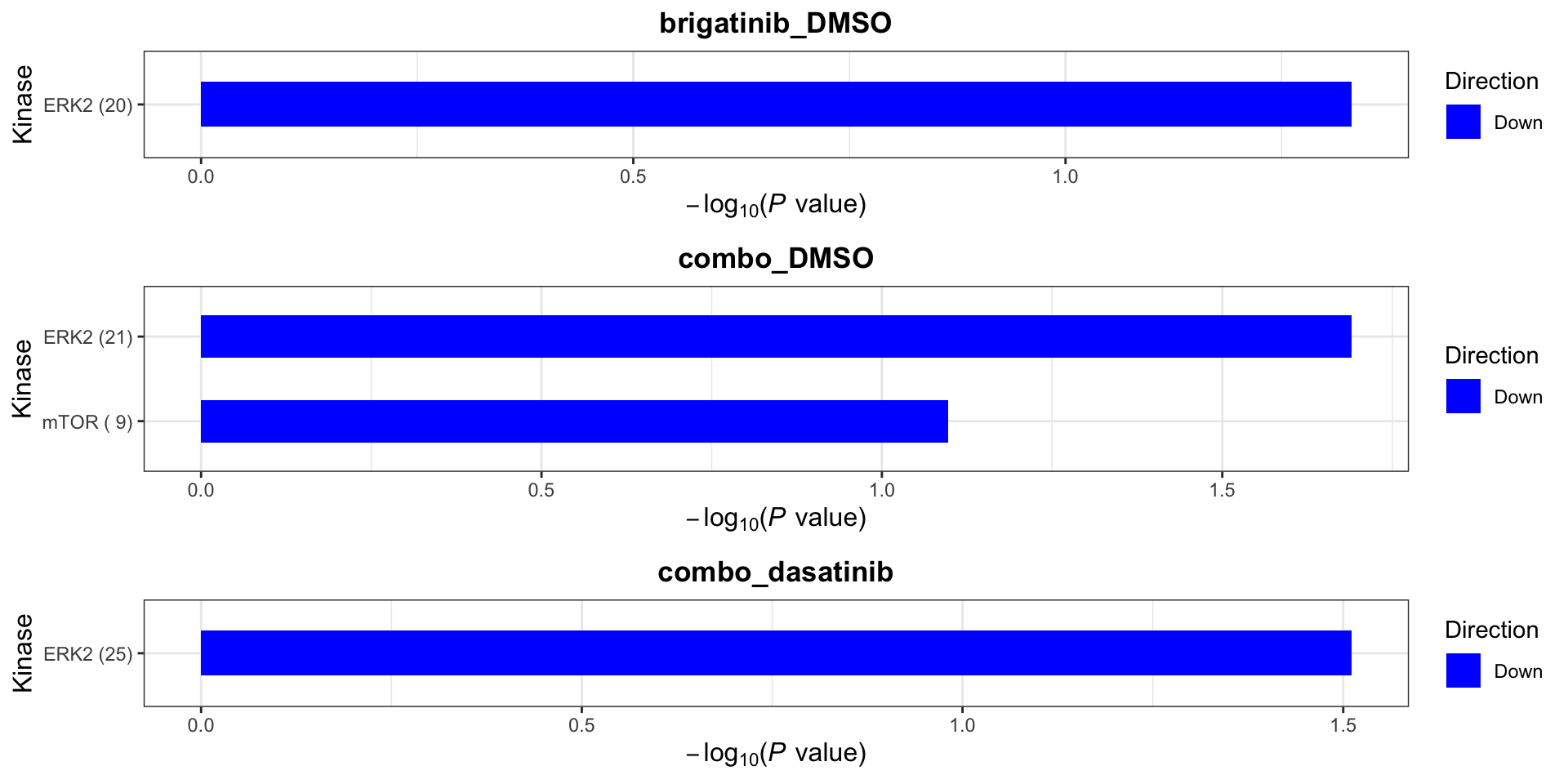
Site-specific Kinase enrichment
Here the site-specificity is considered.
P=0.01 as cut-off
resTab.site <- mutate(resTab, symbol = site)
plotList <- runGeneSetEnrichment(resTab.site, kins, genePCut = 0.05, pCutSet = 0.01, setFdr = FALSE, method="fisher")[1] "Testing for: Kinase"
[1] "Condition: dasatinib_DMSO"
[1] "Condition: brigatinib_DMSO"
[1] "Condition: combo_DMSO"
[1] "Condition: combo_brigatinib"
[1] "Condition: combo_dasatinib"
[1] "No sets passed the criteria"
[1] "No sets passed the criteria"plotList$Kinase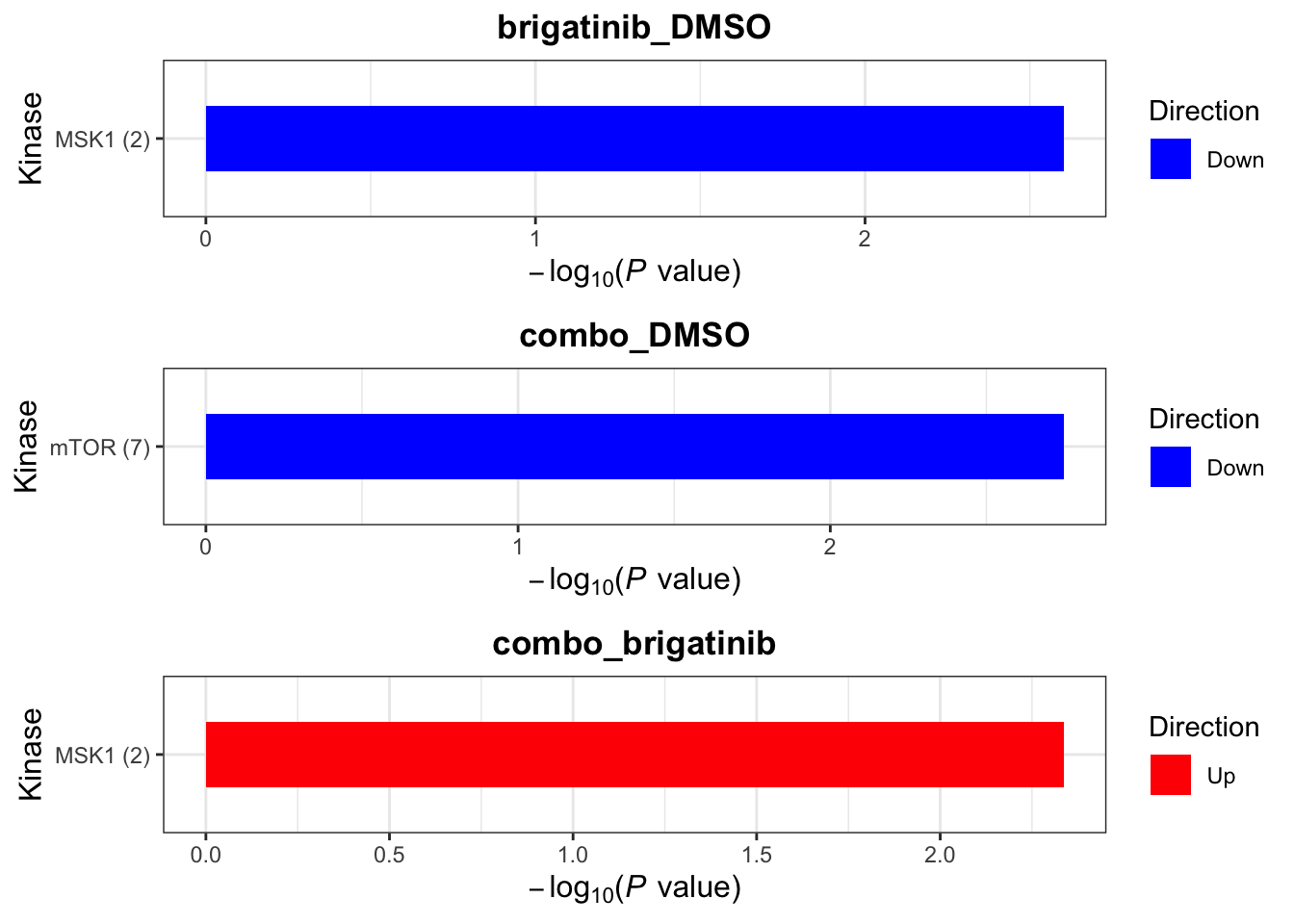
Focuse on the combination effect between Brigatinib and Dasatinib (Interaction effect)
Since Brigatinib and Dasatinib has synergistic effect on cell survival, this part is to detect the synergistic effect also on protein expression level. In statistical term, this is an interaction effect, where the effect of two variables is beyond symbol linear additive effect.
Differential test
Used save results
resTab <- allResList$diffPhos$time_0.17 %>% filter(compare =="interaction")
resTab.sig <- filter(resTab, adj_pval <=0.1)PCA plot with proteins showing interactions
PCA
PC1 versus PC2
plotPCA(fpeSub[resTab.sig$name,], assayName = "imputed", "PC1", "PC2", label ="replicate")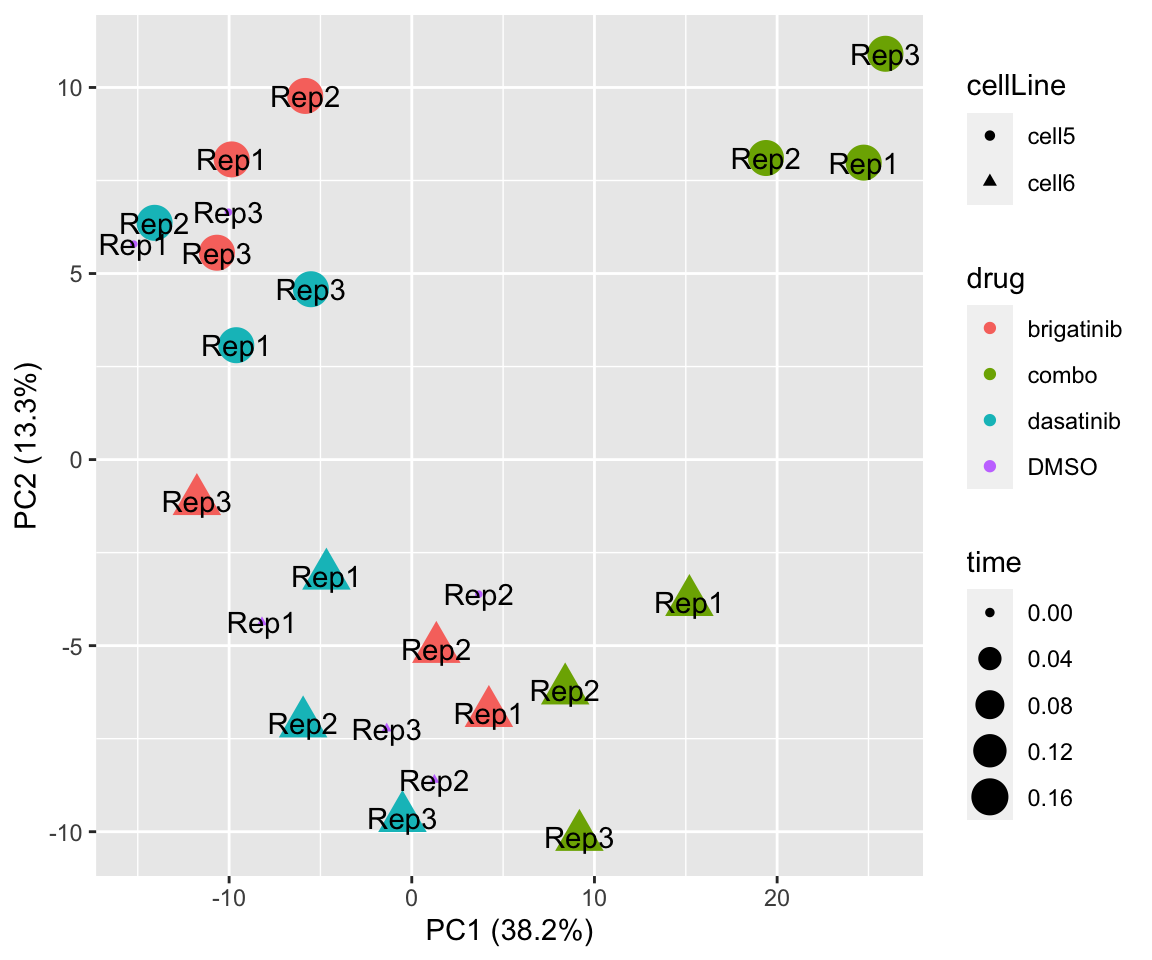 Potential problem with 1 replicate
Potential problem with 1 replicate
PC3 versus PC4
plotPCA(fpeSub[resTab.sig$name,], assayName = "imputed", "PC3", "PC4", label ="replicate")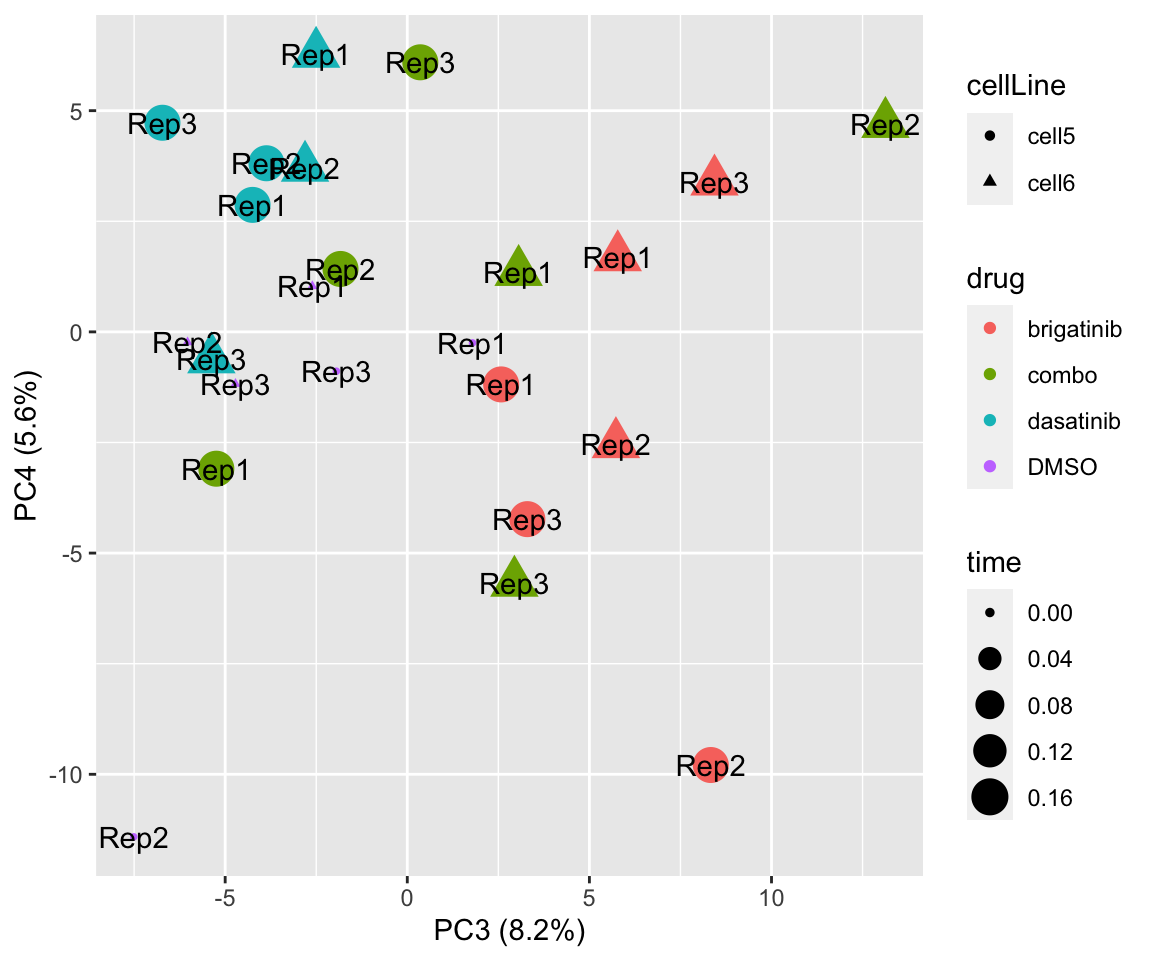
List of proteins with significant interactions (10%FDR)
resTab.sig %>% mutate(across(where(is.numeric), formatC, digits=2)) %>% DT::datatable()Boxplot of top 20 examples
plotTab <- jyluMisc::sumToTidy(fpeSub)
plotList <- lapply(seq(nrow(resTab.sig)), function(i) {
rec <- resTab.sig[i,]
eachTab <- filter(plotTab, rowID == rec$name)
ggplot(eachTab, aes(x=drug, y=Intensity)) +
geom_boxplot(aes(fill = drug)) +
ggbeeswarm::geom_quasirandom() +
facet_wrap(~cellLine) +
theme(legend.position = "none") +
ggtitle(rec$site)
})
cowplot::plot_grid(plotlist = plotList[1:20], ncol=2)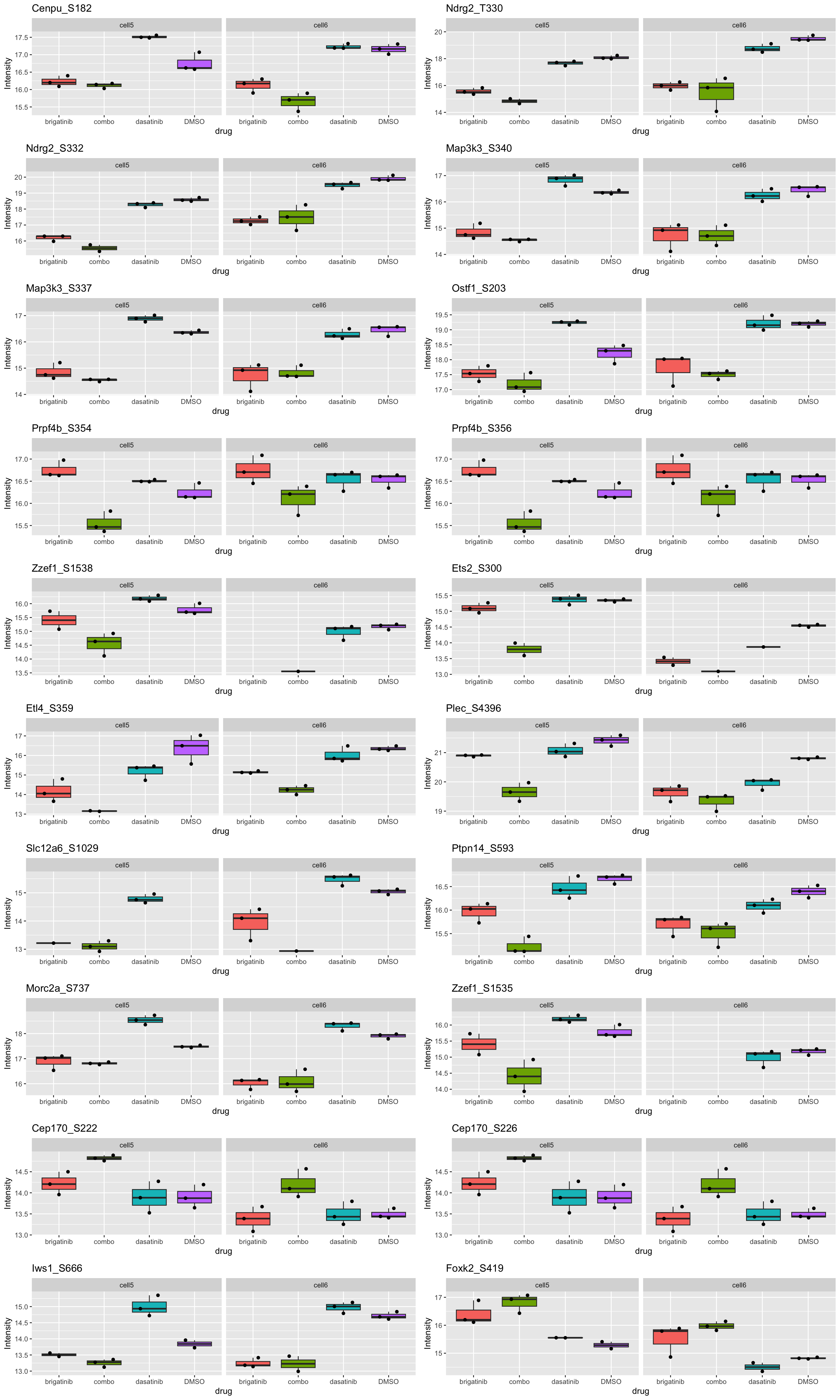
#plot all case in pdf file
#jyluMisc::makepdf(plotList, "../docs/boxplot_interactionPhos_0.17_RUN5.pdf", ncol = 2, nrow =5, height = 15, width = 10)Download the pdf file for all significant interactions: pdf file
Heatmap of top 100 examples
plotProteinHeatmap(resTab, fpeSub, "all", fdrCut = 0.1, ifFDR = TRUE)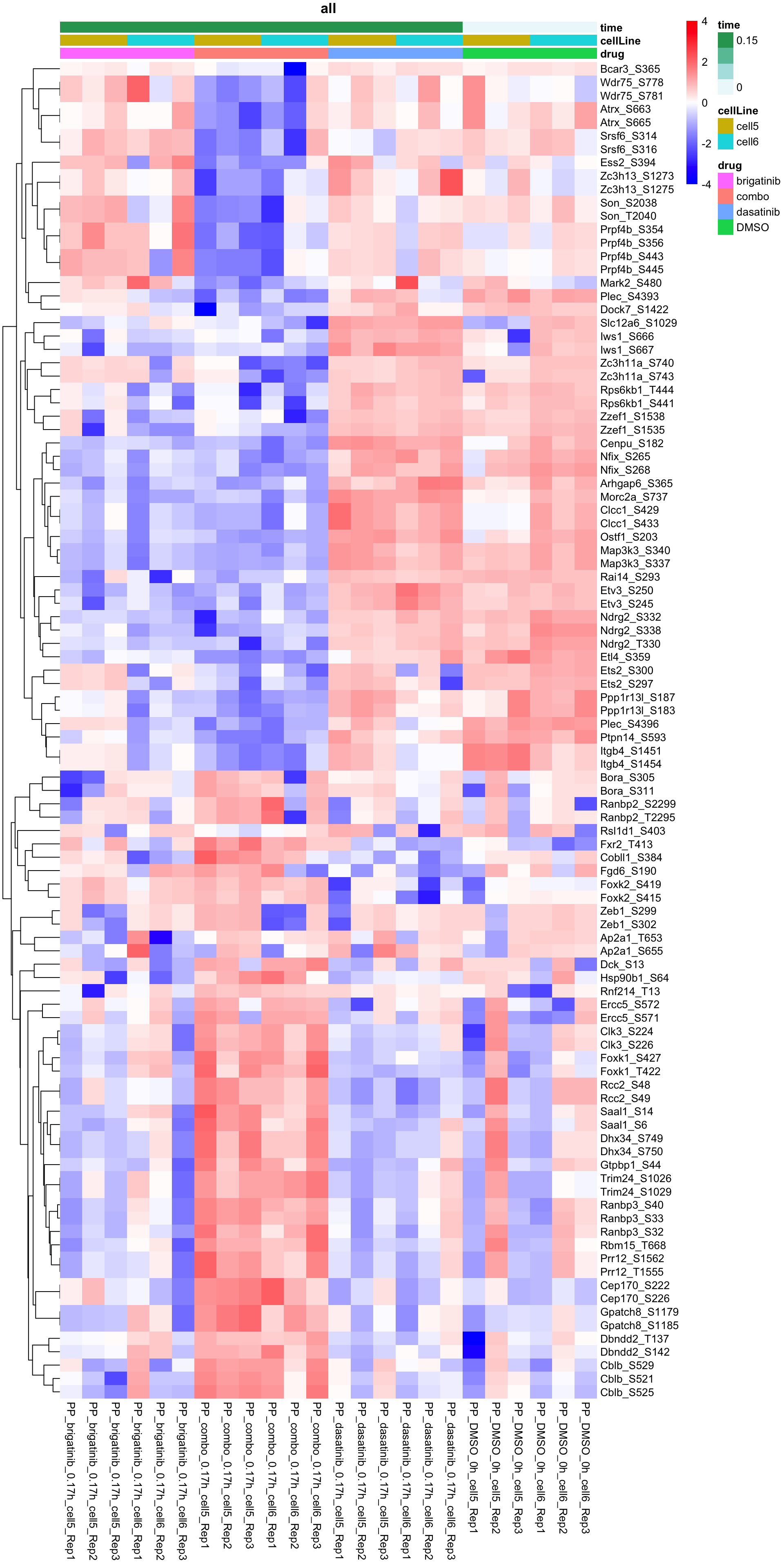
Gene set enrichment analysis
Define useful genesets
plotList <- runGeneSetEnrichment(resTab, gmts, genePCut = 1, pCutSet = 0.05, setFdr = FALSE, method="gsea", collapsePathway = TRUE)[1] "Testing for: Hallmark"
[1] "Condition: interaction"
[1] "Testing for: CanonicalPathway"
[1] "Condition: interaction"
[1] "Testing for: Kinase"
[1] "Condition: interaction"
[1] "No sets passed the criteria"Hallmarks
plotList$Hallmark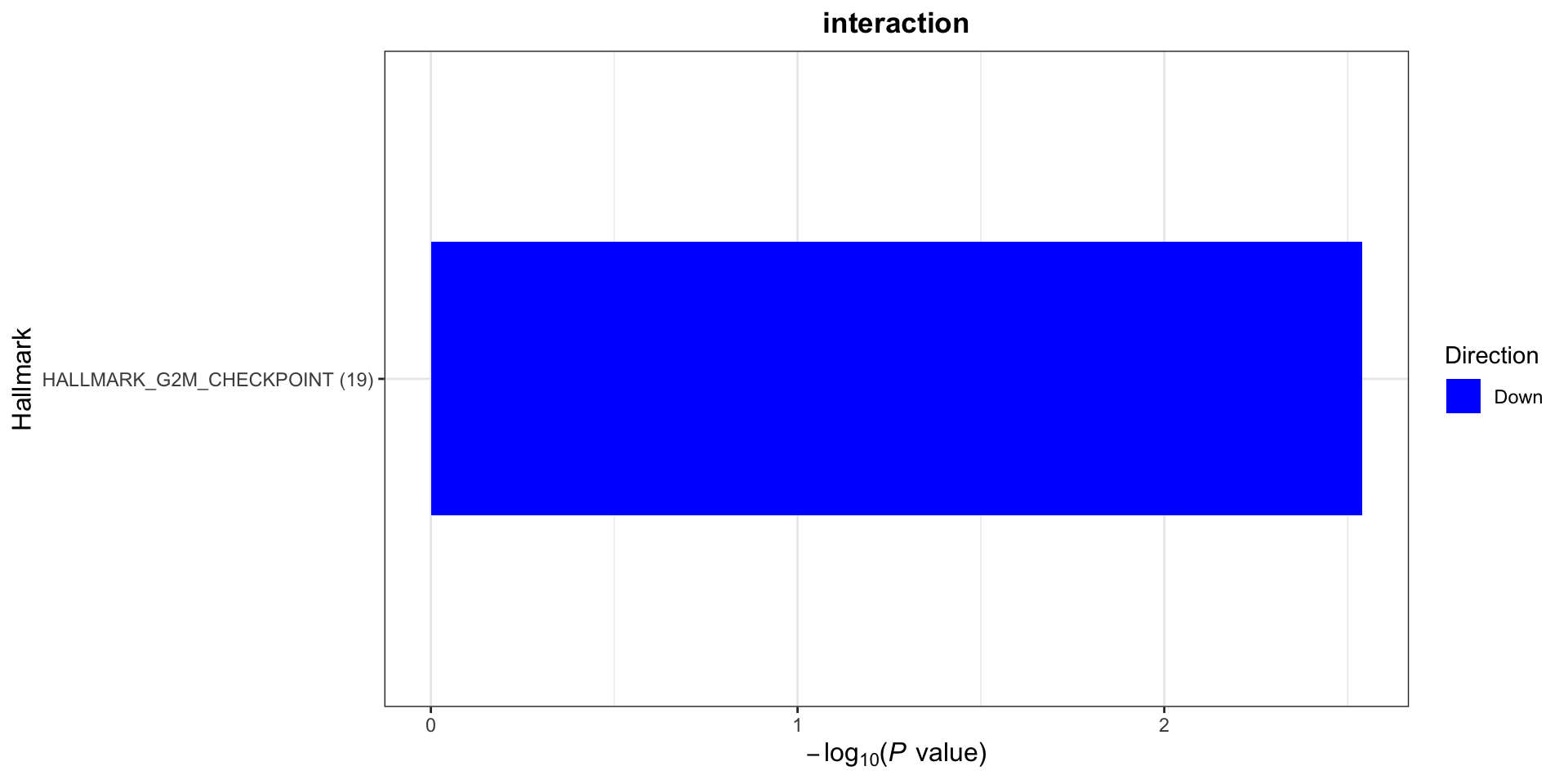
Canonical pathways
plotList$CanonicalPathway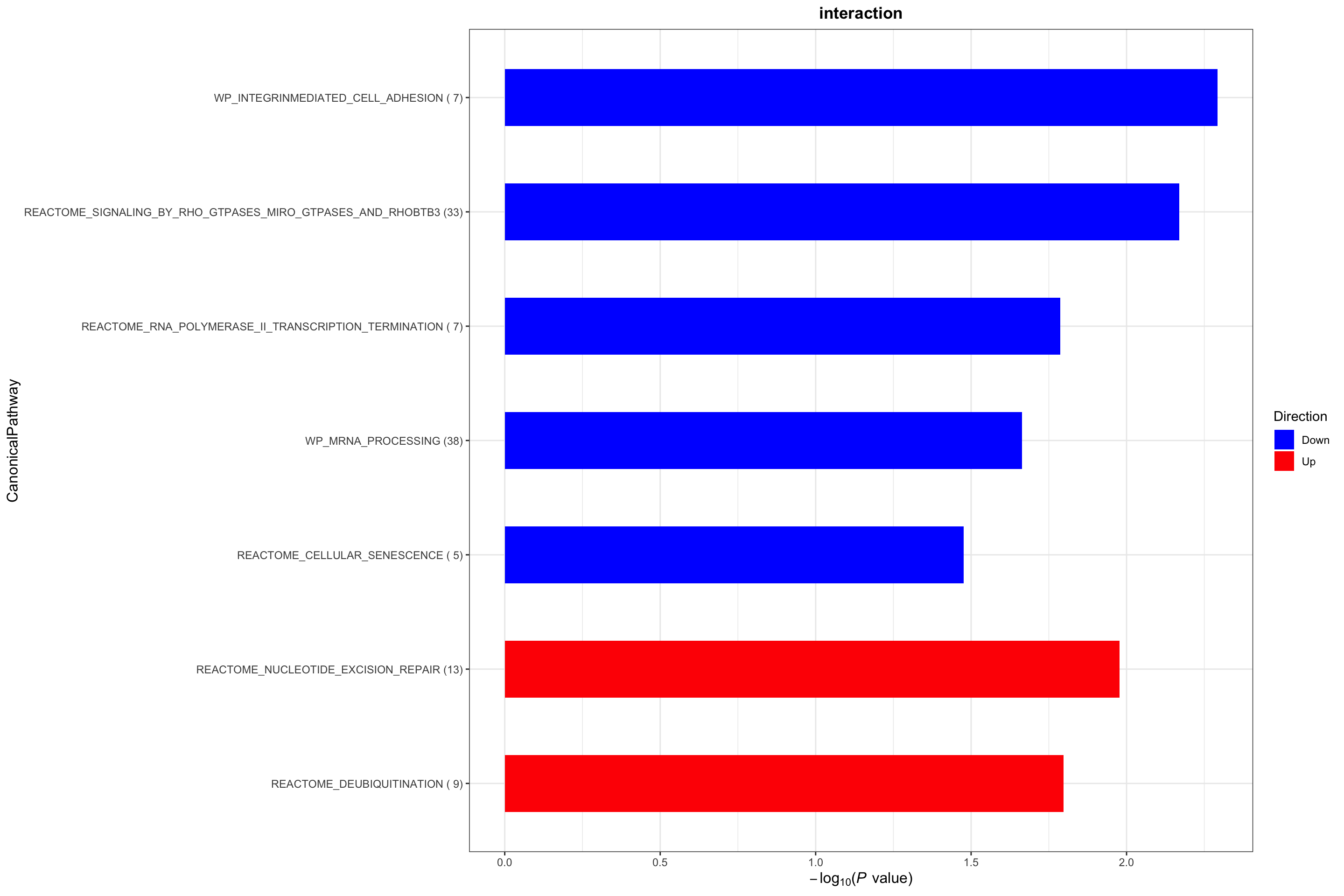
List of leading edge genes for each pathway
Leading edges genes are not necessarily significantly differentially expressed, but they contribute most to the enrichment analysis. Please see explaination of leading edge genes on this page: https://www.gsea-msigdb.org/gsea/doc/GSEAUserGuideFrame.html
DT::datatable(plotList$leadingEdgeGene)writexl::write_xlsx(plotList$leadingEdgeGene, path = "../docs/enrichment_tables/phos_gsea_0.16combo_all.xlsx")Site-unspecific Kinase enrichment
On protein level, site specificity is not considered. This is because many phosphorylation sites lack the up-stream kinase annotation in the database. We can use a less stringent criteria to define kinase-substrate relation ship
plotList$Kinase[1] "No sets passed the criteria"Site-specific Kinase enrichment
Here the site-specificity is considered.
P=0.01 as cut-off
resTab.site <- mutate(resTab, symbol = site)
plotList <- runGeneSetEnrichment(resTab.site, kins, genePCut = 0.05, pCutSet = 0.01, setFdr = FALSE, method="fisher")[1] "Testing for: Kinase"
[1] "Condition: interaction"
[1] "No sets passed the criteria"plotList$Kinase[1] "No sets passed the criteria"Plot interesting pathways
BIOCARTA_MAPK_PATHWAY
plotSetHeatmap(fpeSub, filter(resTab, pval < 0.05), drugPair = "all", gmtFile = gmts$CanonicalPathway,
setName = "BIOCARTA_MAPK_PATHWAY")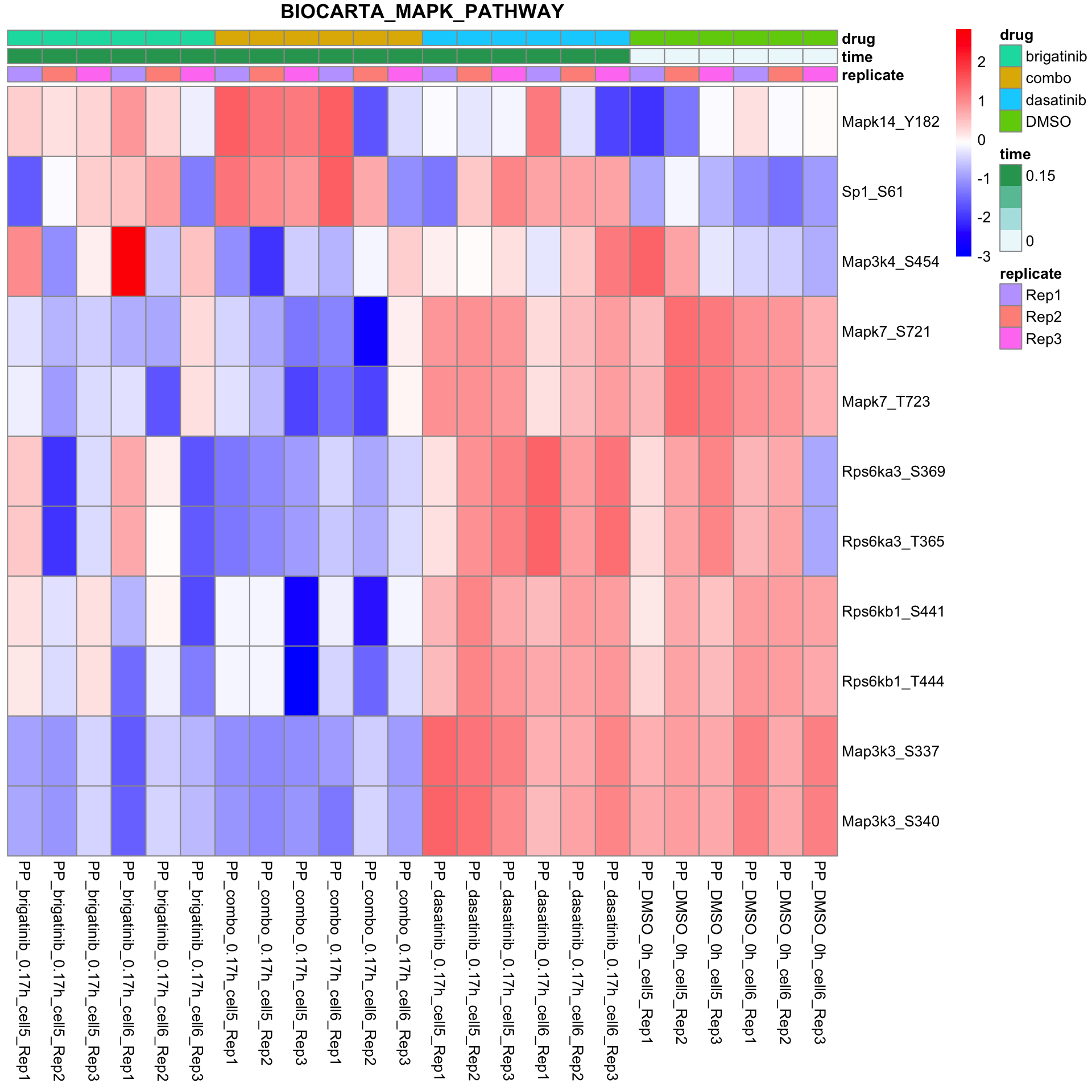
G2M_CHECKPOINT
All conditions will be included, difference among cell lines will be regressed out for better visualization
plotSetHeatmap(fpeSub, filter(resTab, pval < 0.05), drugPair = "all", gmtFile = gmts$Hallmark,
setName = "HALLMARK_G2M_CHECKPOINT")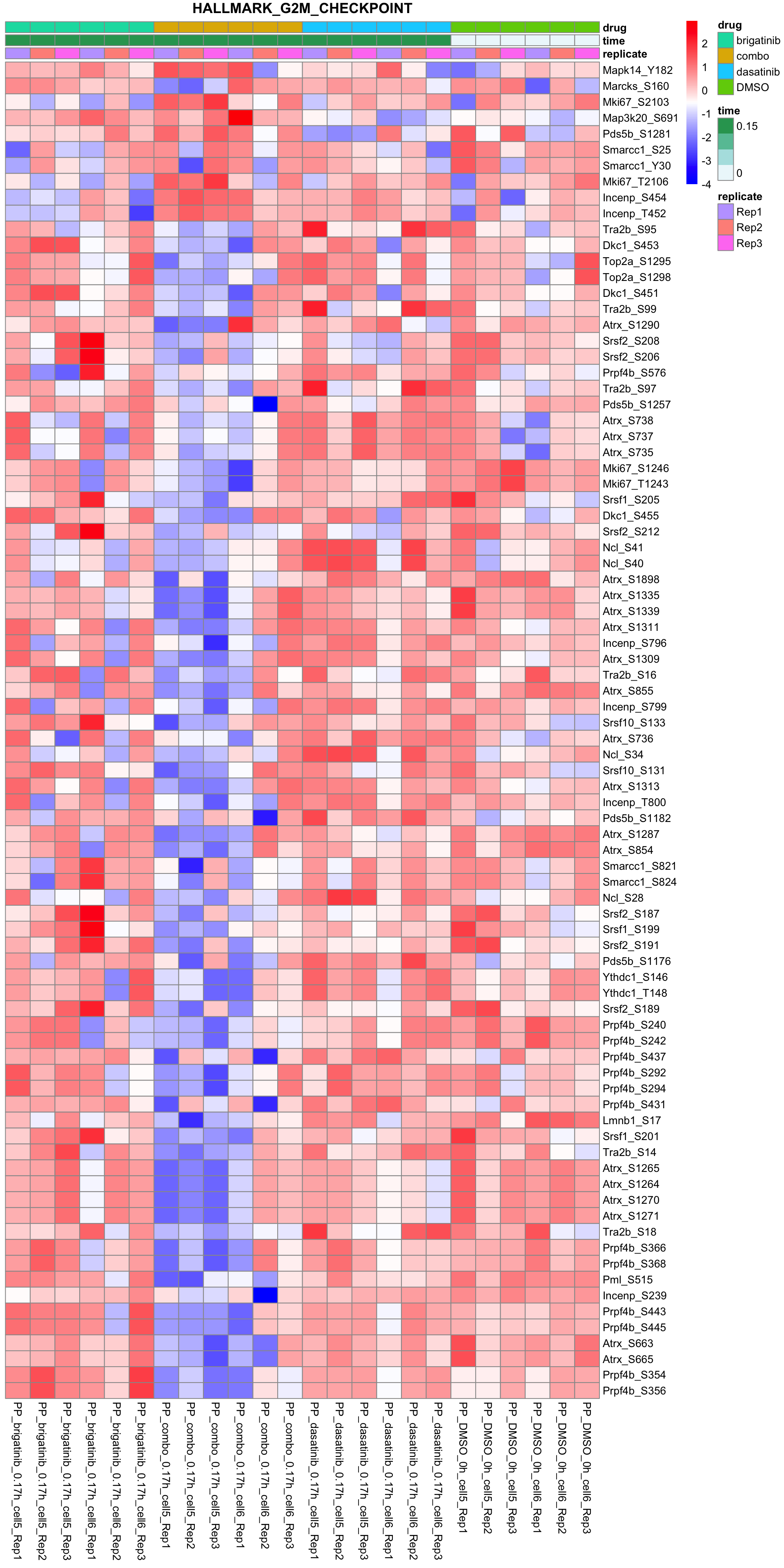
Differential expression at 16 hours for each drug
Preprocessing
filterList <- list(time = c(16,0))
fpeSub <- preprocessPhos(maeData, filterList, missCut = 0.5, transform = "vst", normalize = TRUE)[1] "Number of proteins and samples:"
[1] 4247 24Plot value distribution
plotTab <- jyluMisc::sumToTidy(fpeSub)
ggplot(plotTab, aes(x=sample, y=Intensity)) +
geom_boxplot(aes(fill = drug)) +
theme(axis.text.x = element_text(angle = 90, hjust = 1, vjust = 0.5))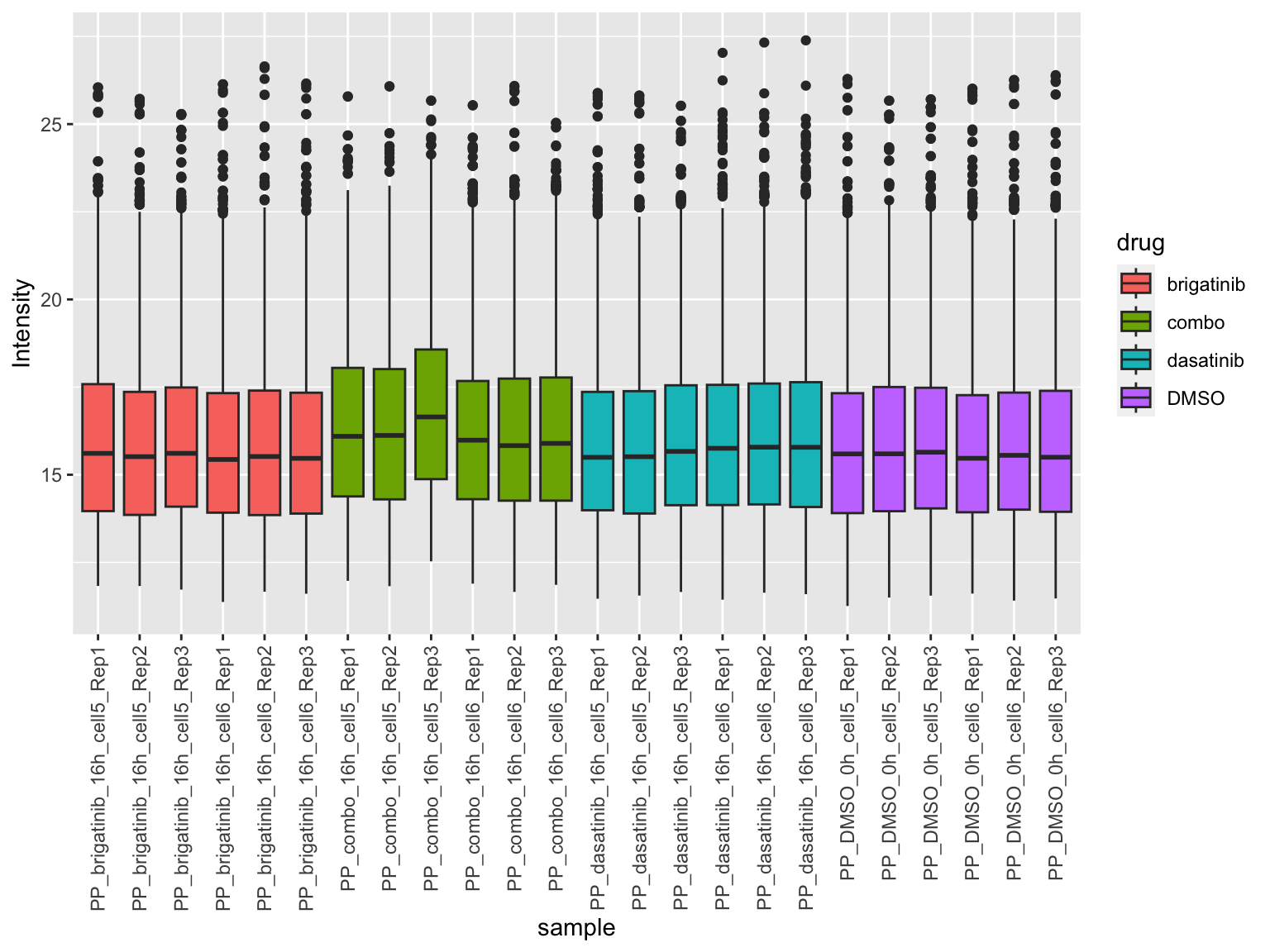
PCA
PC1 versus PC2
plotPCA(fpeSub, assayName = "imputed", "PC1", "PC2", topVar = 2000, label ="replicate")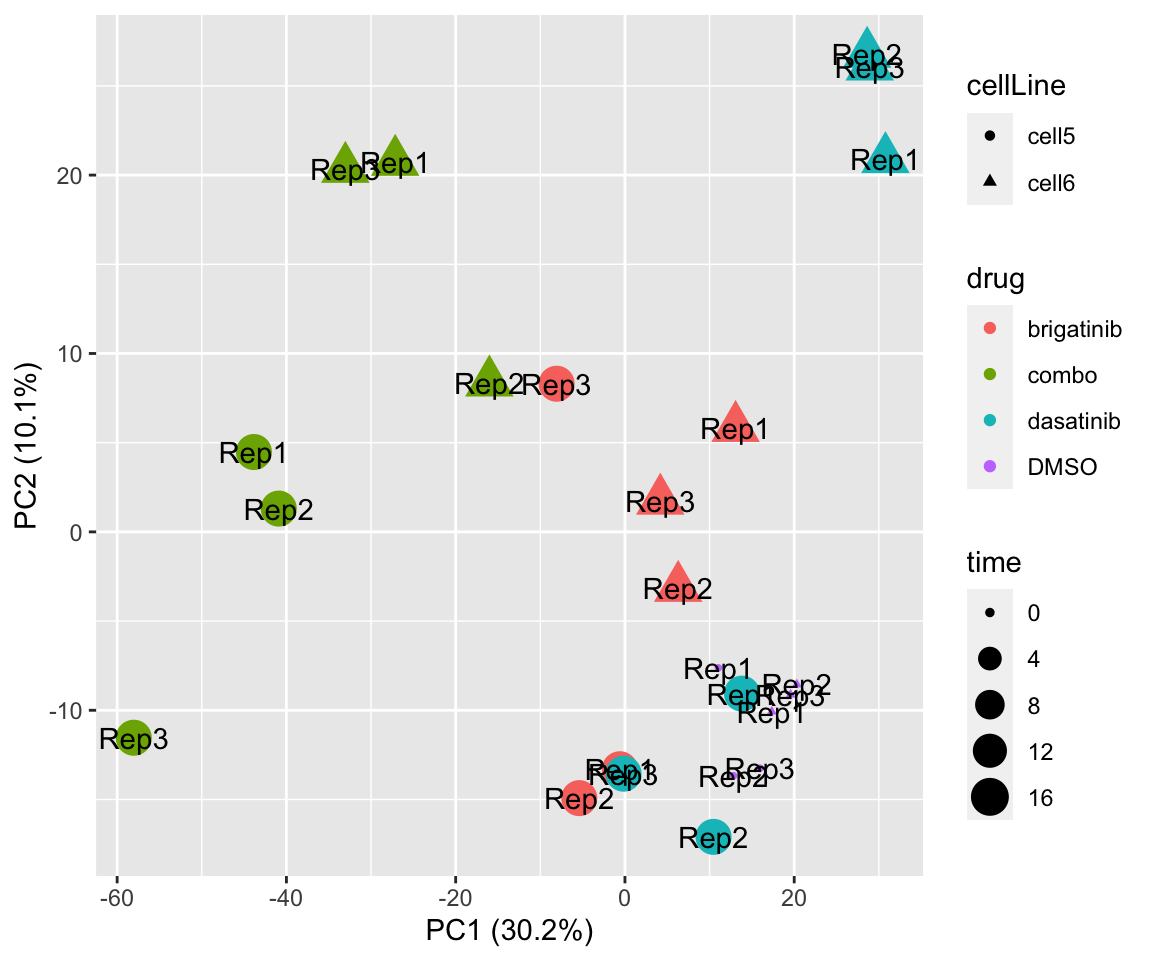 Potential problem with 1 replicate
Potential problem with 1 replicate
PC3 versus PC4
plotPCA(fpeSub, assayName = "imputed", "PC3", "PC4", topVar = 2000, label = "replicate")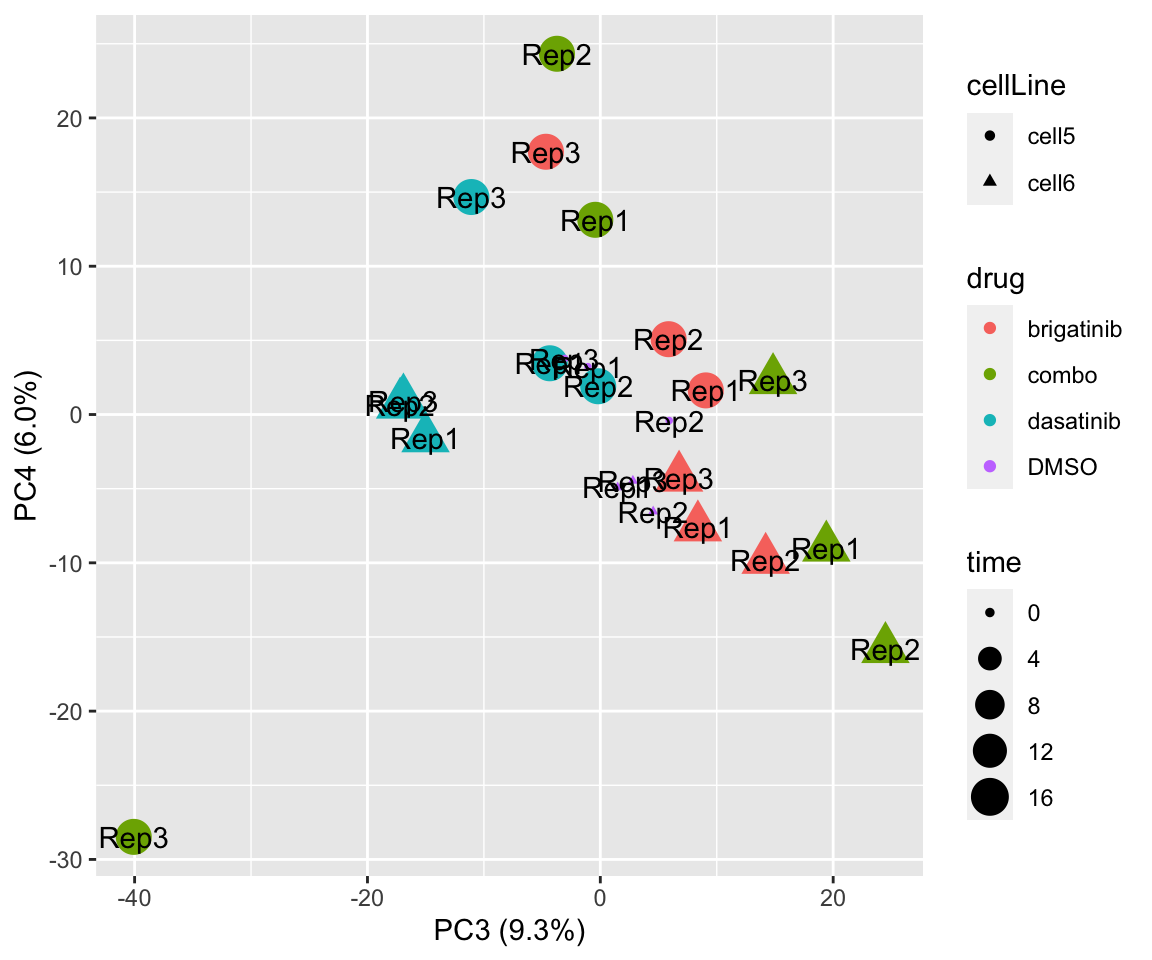
Differential expression using proDA
Used save results
resTab <- allResList$diffPhos$time_16 %>% filter(compare !="interaction")Table of significant associations (1% FDR)
Here I used 1% FDR, otherwise there are too many proteins
resTab.sig <- filter(resTab, adj_pval <= 0.05)
resTab.sig %>% mutate(across(where(is.numeric), formatC, digits=2)) %>% DT::datatable()P-value histogram for each comparison
ggplot(resTab, aes(x=pval)) +
geom_histogram(bins = 20, fill = "lightblue", color = "grey50") +
facet_wrap(~compare)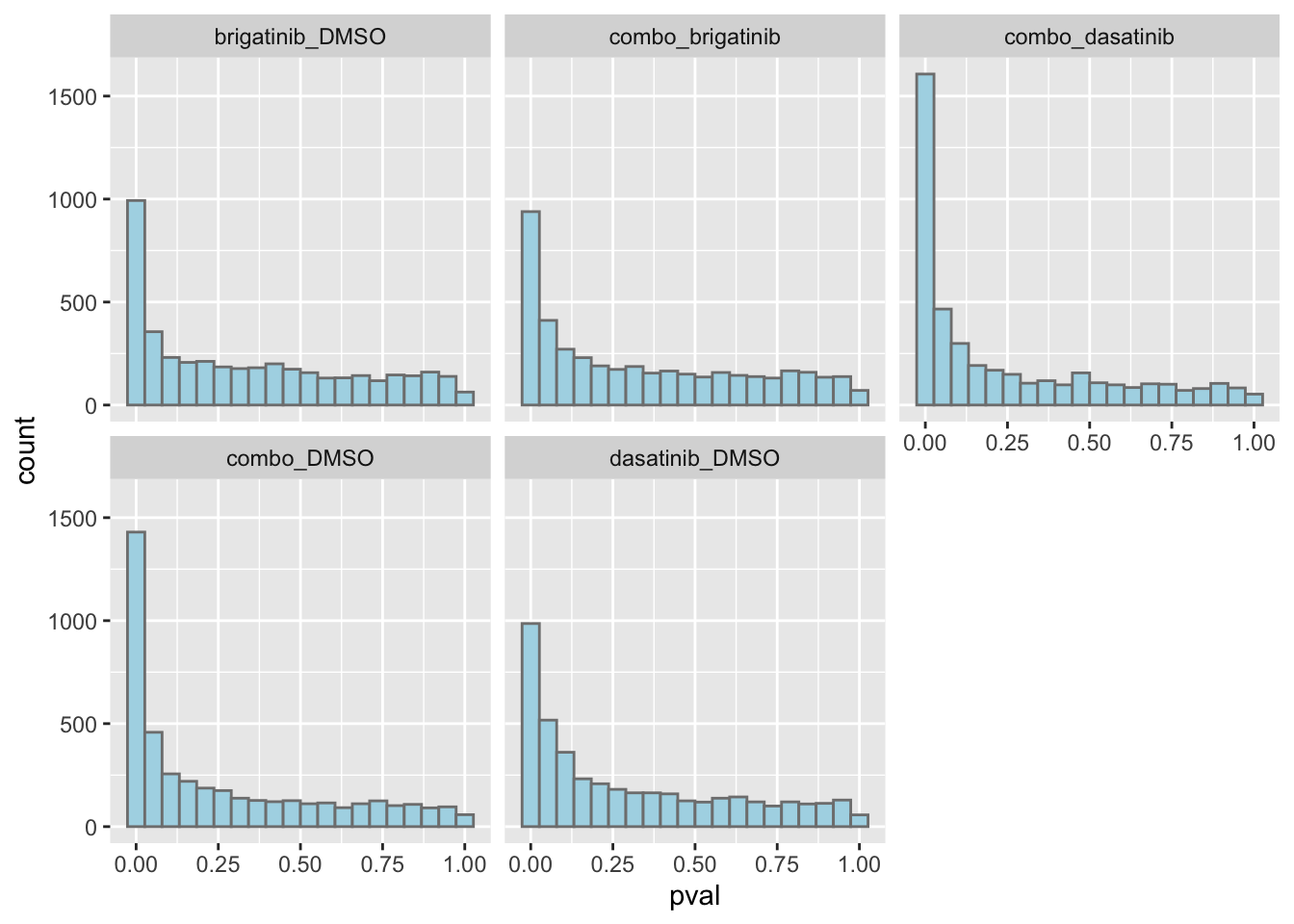
Number of DE proteins for each comparison (5% FDR)
sumTab <- filter(resTab, adj_pval < 0.05) %>%
mutate(direction = ifelse(diff>0, "Up","Down")) %>%
group_by(compare, direction) %>%
summarise(n=length(name))
ggplot(sumTab, aes(x=compare, y=n)) +
geom_bar(aes(fill = direction), stat = "identity", position = "dodge") +
coord_flip() +
geom_text(aes(label =n)) +
facet_wrap(~direction, ncol=2, scales = "free_x") +
xlab("Number")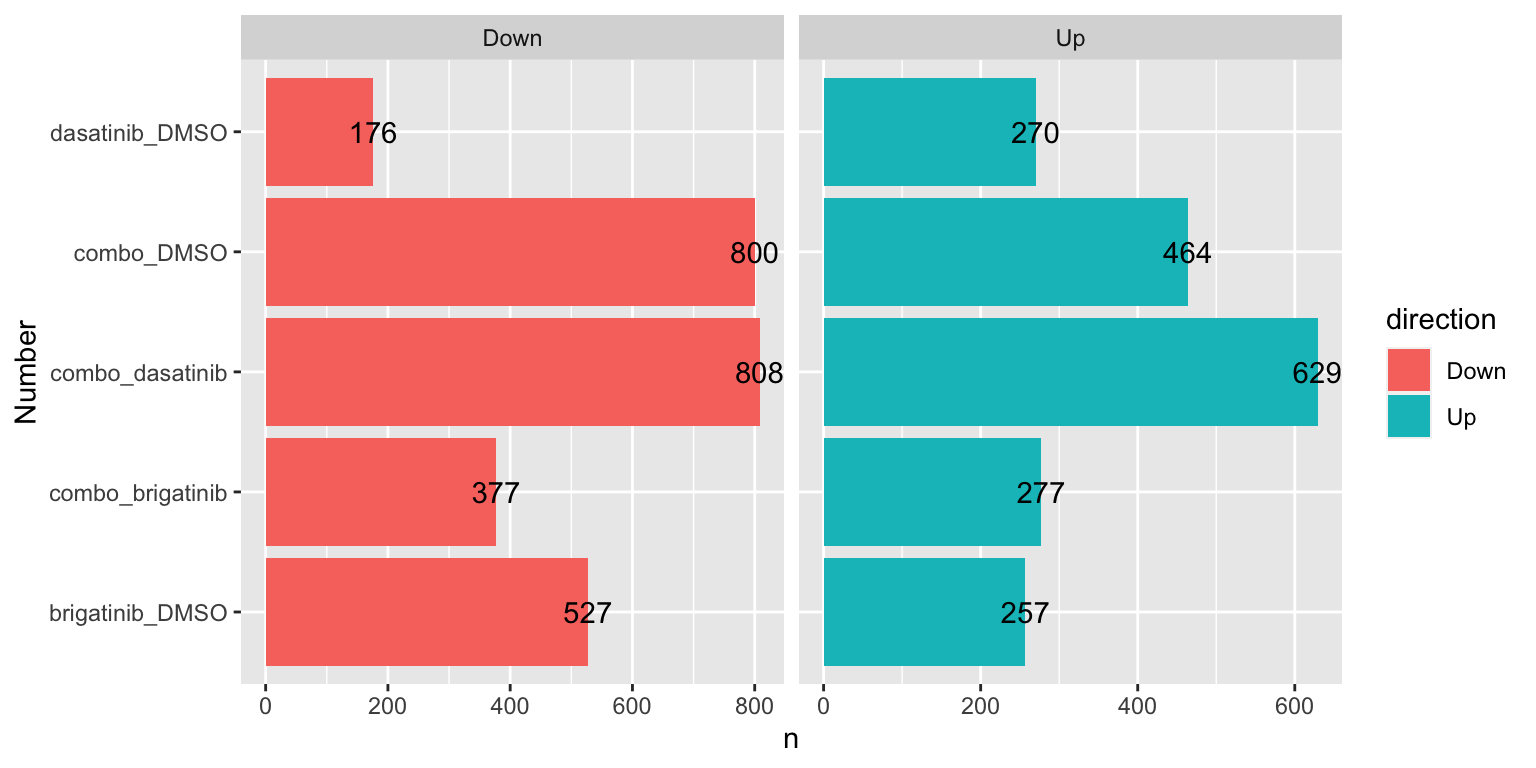
Heatmap of DE proteins for each treatment
Brigatinib versus DMSO
plotProteinHeatmap(resTab, fpeSub, "brigatinib_DMSO", fdrCut = 0.05, ifFDR = TRUE)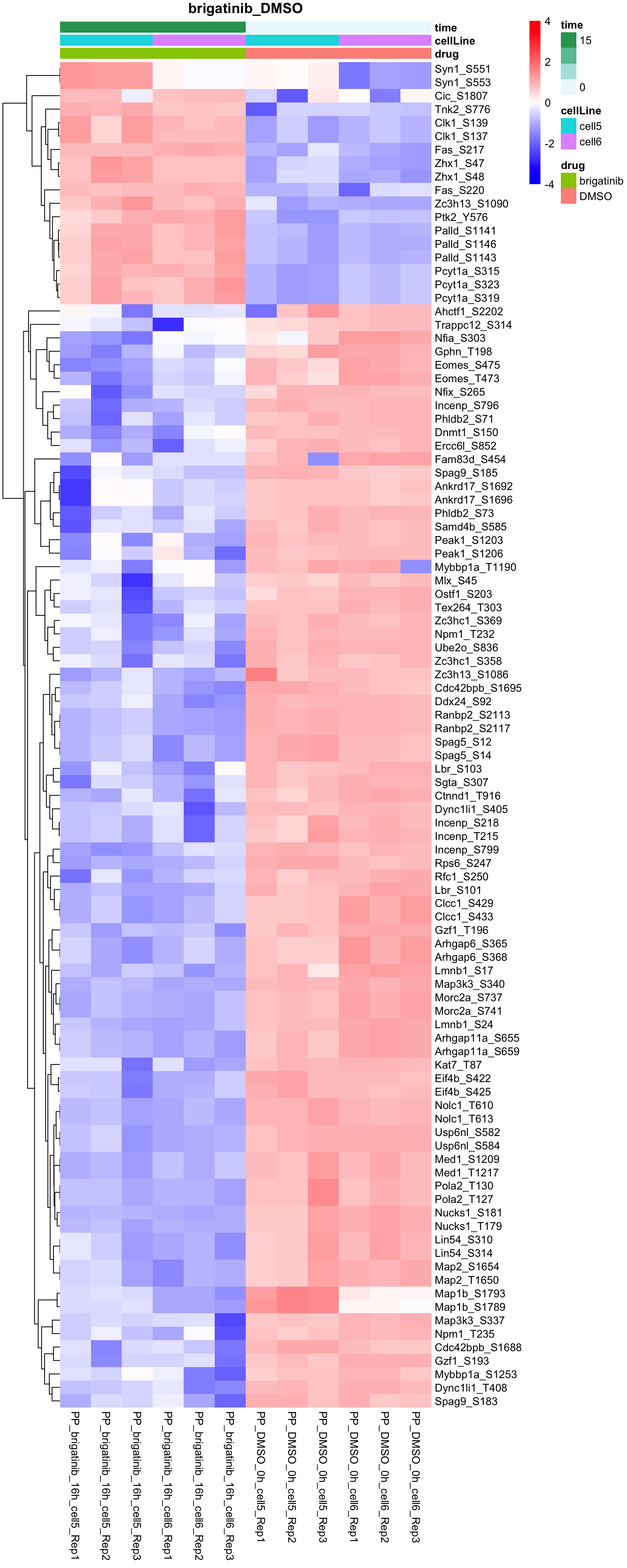
Dasatinib versus DMSO
plotProteinHeatmap(resTab, fpeSub, "dasatinib_DMSO", fdrCut = 0.05, ifFDR =TRUE)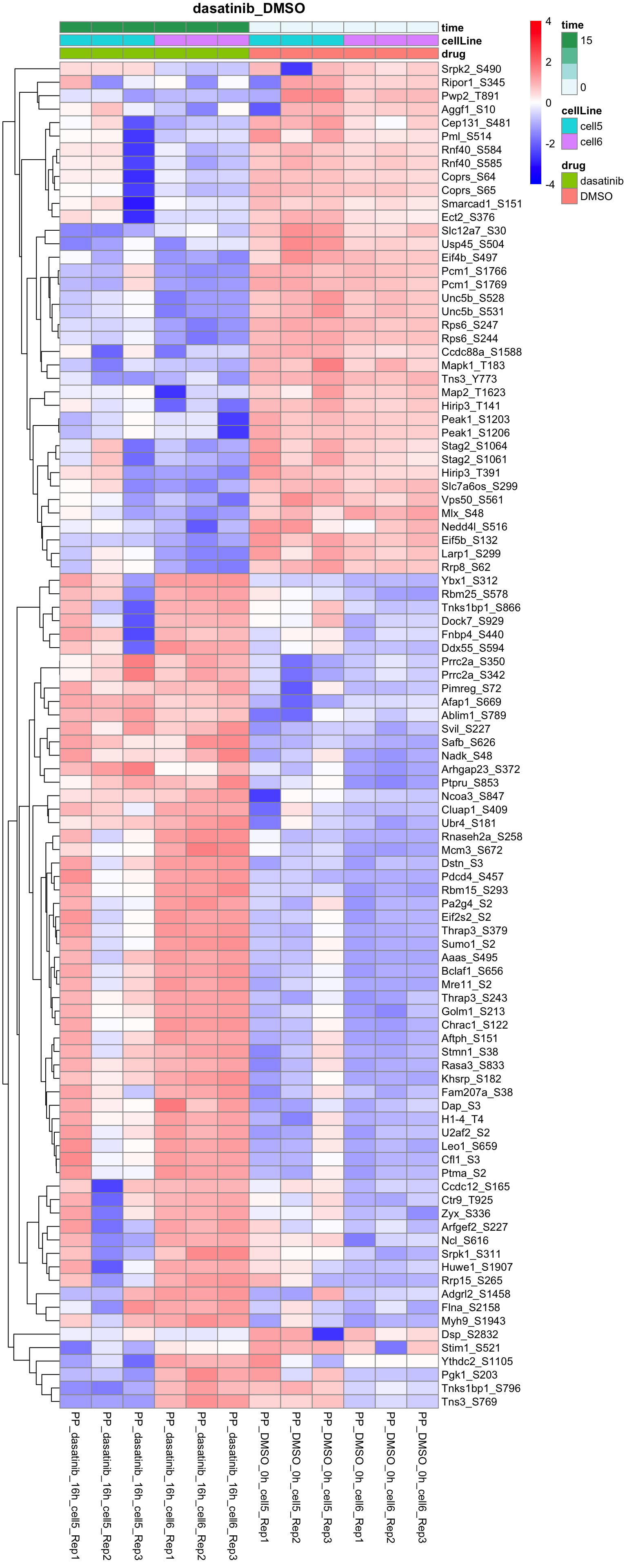
Combo versus DMSO
plotProteinHeatmap(resTab, fpeSub, "combo_DMSO", fdrCut = 0.05, ifFDR = TRUE)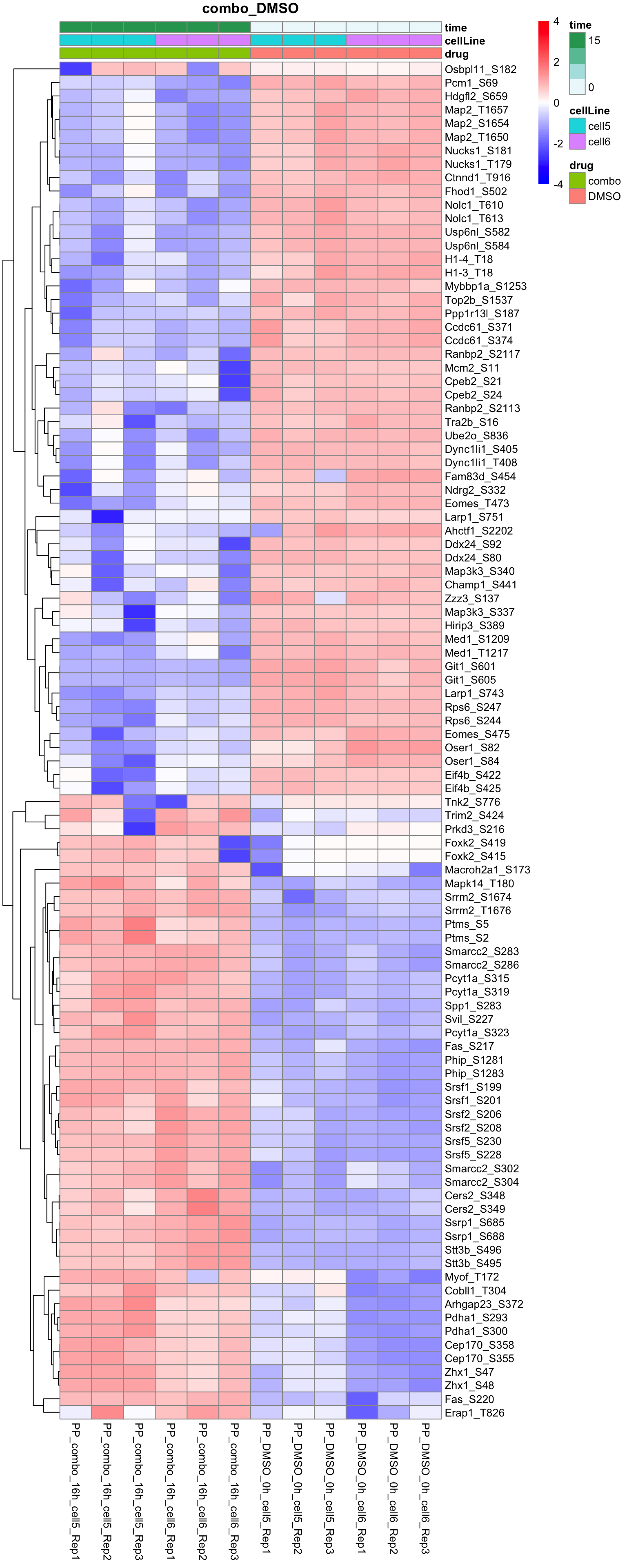
Combo versus brigatinib
plotProteinHeatmap(resTab, fpeSub, "combo_brigatinib", fdrCut = 0.05, ifFDR = TRUE)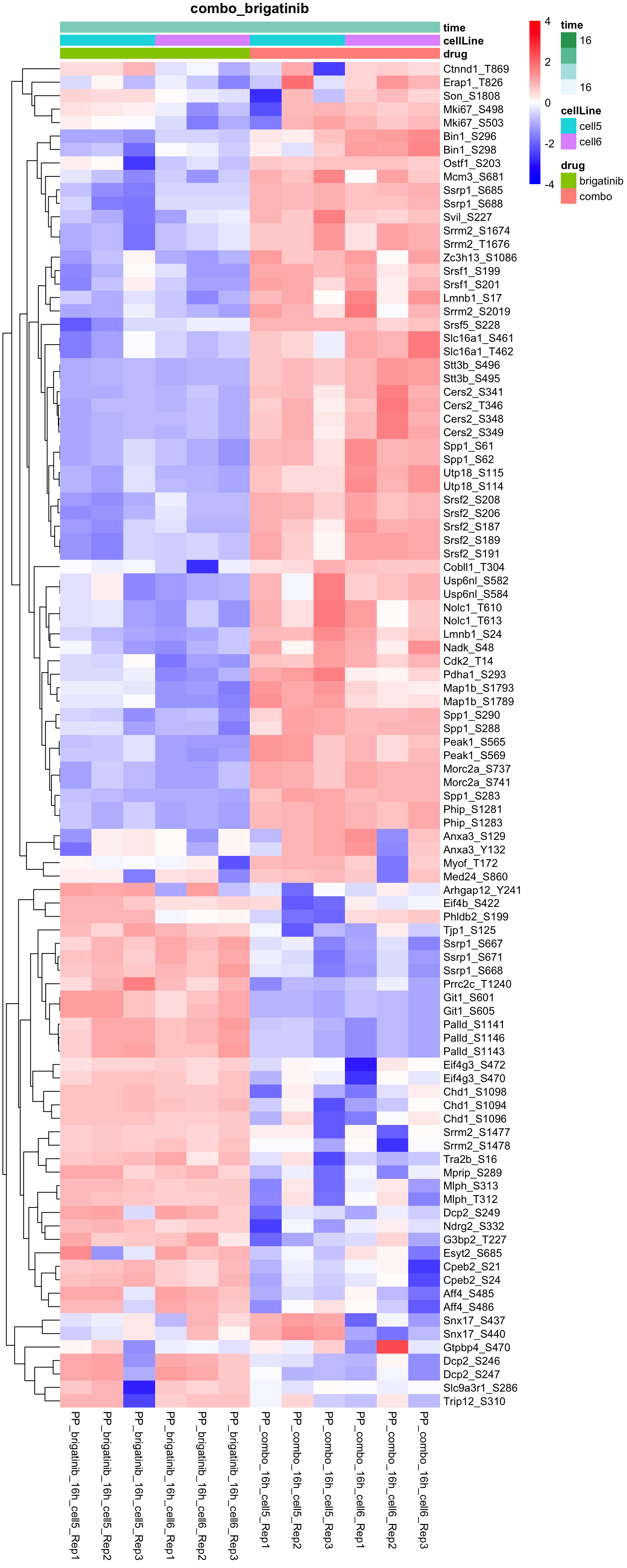
Combo versus dasatinib
plotProteinHeatmap(resTab, fpeSub, "combo_dasatinib", fdrCut = 0.05, ifFDR = TRUE)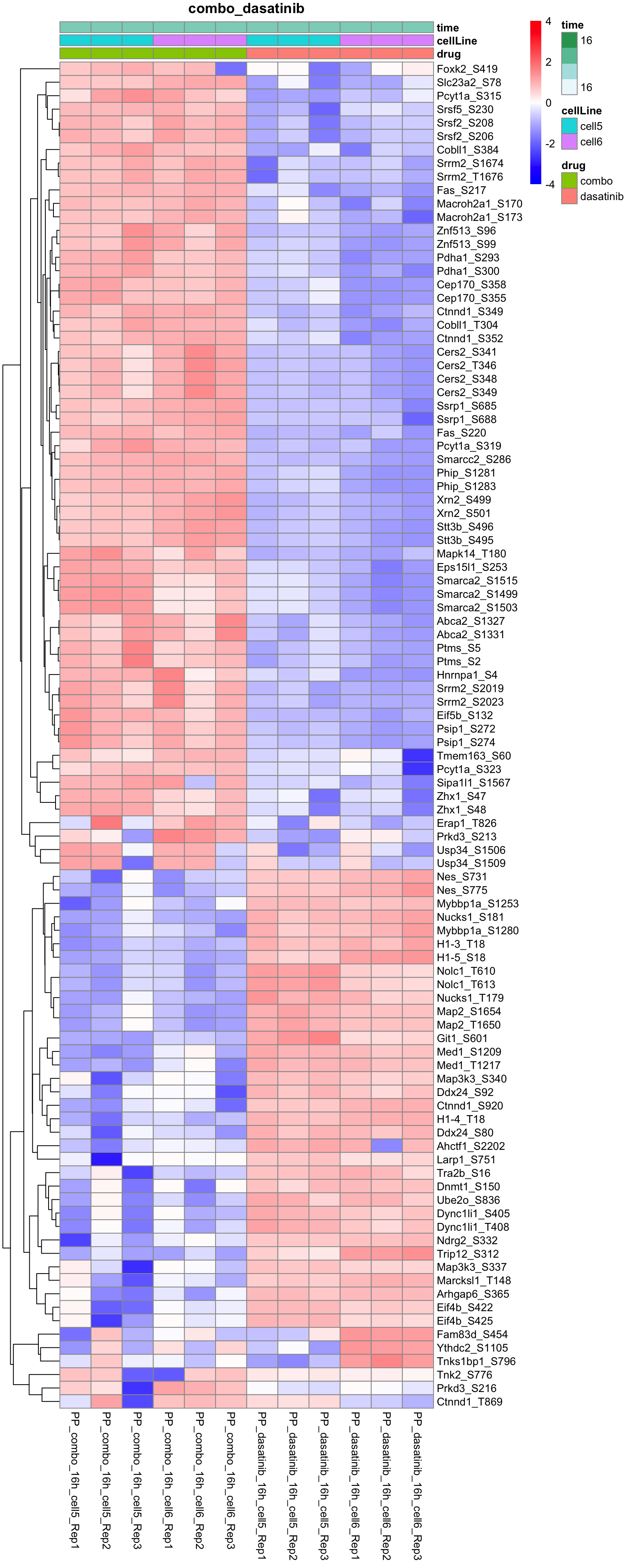
Overlap of DE proteins
Up regulated
deList <- lapply(unique(resTab$compare), function(x) {
filter(resTab, compare == x,adj_pval<0.1, diff >0)$name
})
names(deList) <- unique(resTab$compare)
upset(fromList(deList))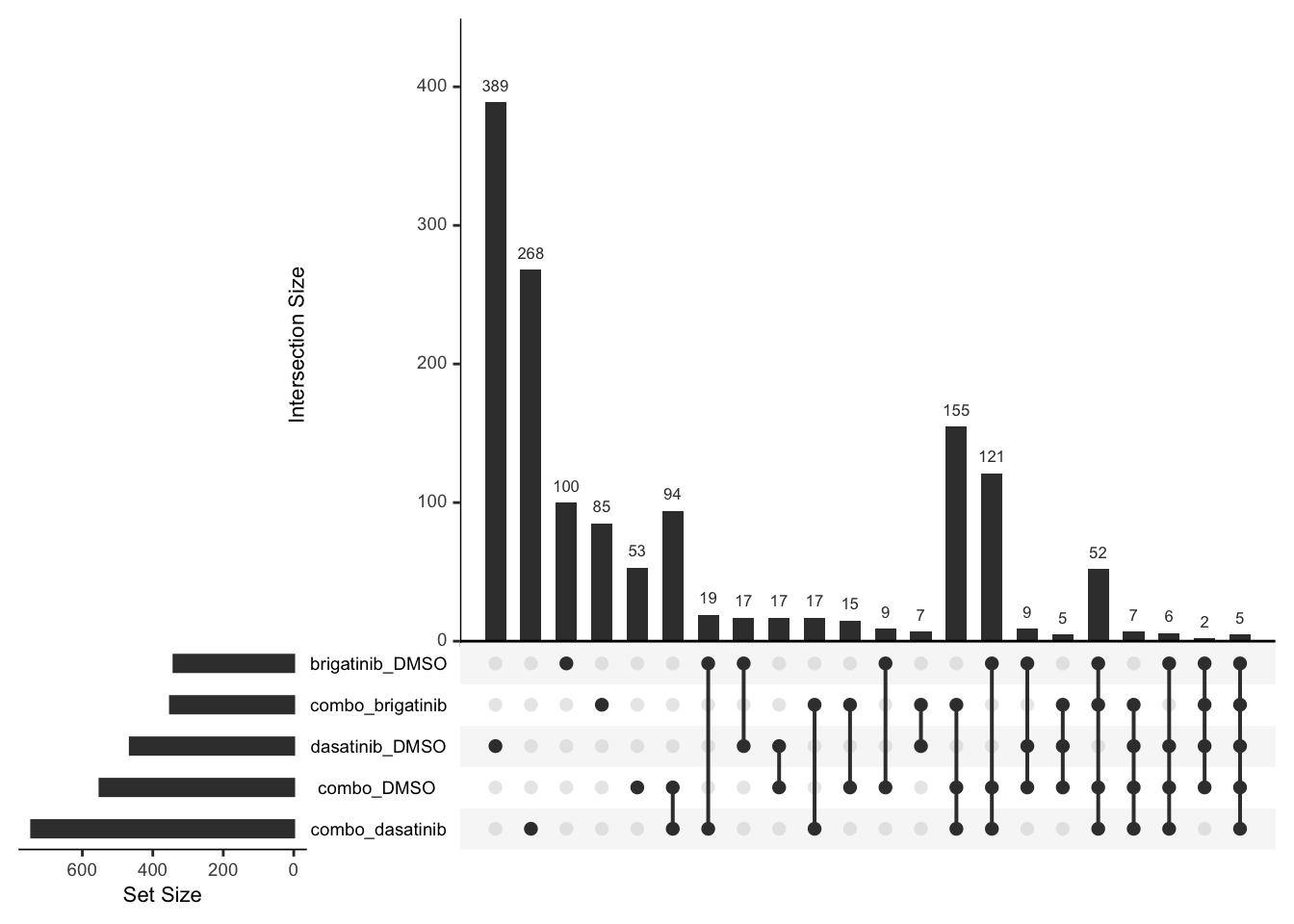
Down regulated
deList <- lapply(unique(resTab$compare), function(x) {
filter(resTab, compare == x, adj_pval <= 0.1, diff <0)$name
})
names(deList) <- unique(resTab$compare)
upset(fromList(deList))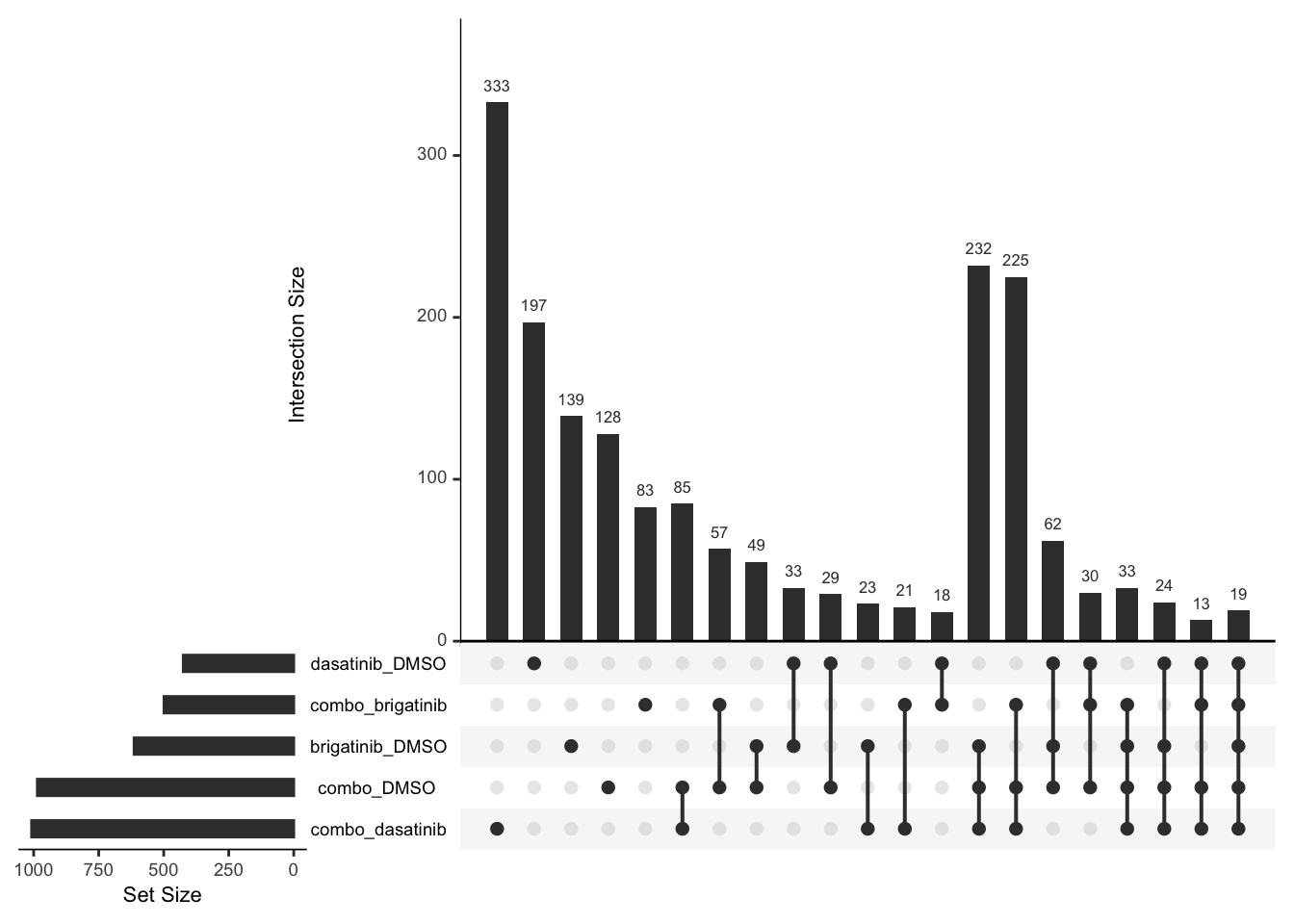
Gene set enrichment analysis
Define useful genesets
plotList <- runGeneSetEnrichment(resTab, gmts, genePCut = 1, pCutSet = 0.05, geneFdr = FALSE, setFdr = TRUE, method="gsea", collapsePathway = TRUE)[1] "Testing for: Hallmark"
[1] "Condition: dasatinib_DMSO"
[1] "Condition: brigatinib_DMSO"
[1] "Condition: combo_DMSO"
[1] "Condition: combo_brigatinib"
[1] "Condition: combo_dasatinib"
[1] "No sets passed the criteria"
[1] "Testing for: CanonicalPathway"
[1] "Condition: dasatinib_DMSO"
[1] "Condition: brigatinib_DMSO"
[1] "Condition: combo_DMSO"
[1] "Condition: combo_brigatinib"
[1] "Condition: combo_dasatinib"
[1] "No sets passed the criteria"
[1] "No sets passed the criteria"
[1] "No sets passed the criteria"
[1] "No sets passed the criteria"
[1] "Testing for: Kinase"
[1] "Condition: dasatinib_DMSO"
[1] "Condition: brigatinib_DMSO"
[1] "Condition: combo_DMSO"
[1] "Condition: combo_brigatinib"
[1] "Condition: combo_dasatinib"
[1] "No sets passed the criteria"
[1] "No sets passed the criteria"Hallmarks
plotList$Hallmark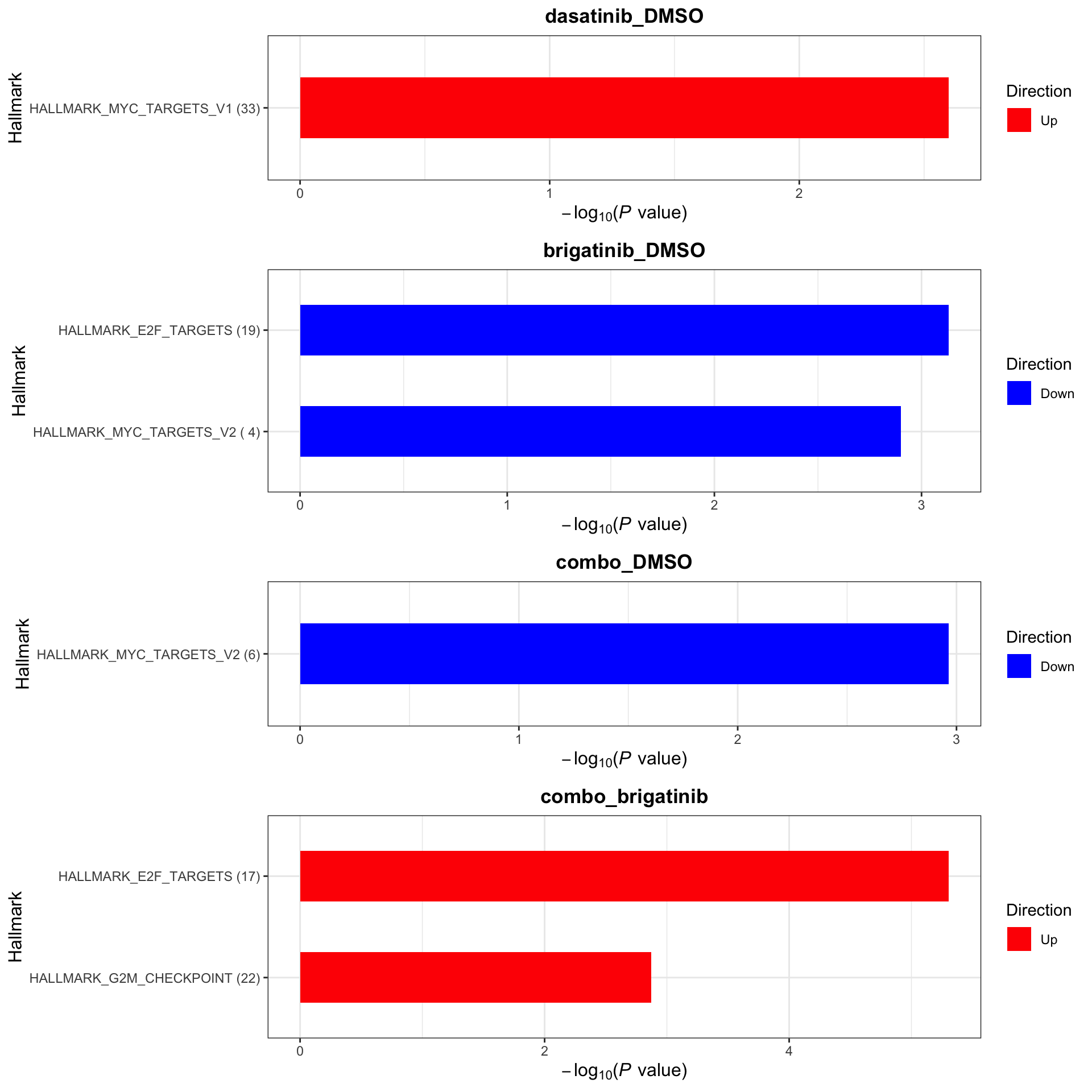
Canonical pathways
plotList$CanonicalPathway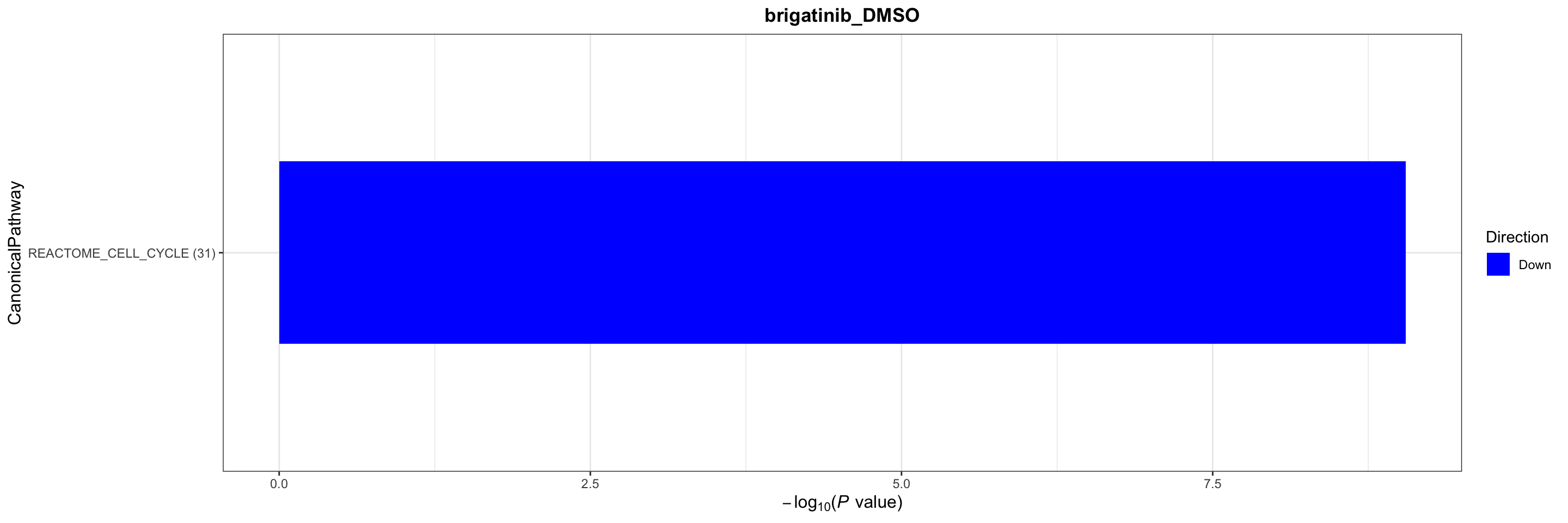
List of leading edge genes for each pathway
Leading edges genes are not necessarily significantly differentially expressed, but they contribute most to the enrichment analysis. Please see explaination of leading edge genes on this page: https://www.gsea-msigdb.org/gsea/doc/GSEAUserGuideFrame.html
DT::datatable(plotList$leadingEdgeGene)writexl::write_xlsx(plotList$leadingEdgeGene, path = "../docs/enrichment_tables/phos_gsea_16_all.xlsx")Site-unspecific Kinase enrichment
On protein level, site specificity is not considered. This is because many phosphorylation sites lack the up-stream kinase annotation in the database. We can use a less stringent criteria to define kinase-substrate relation ship
plotList$Kinase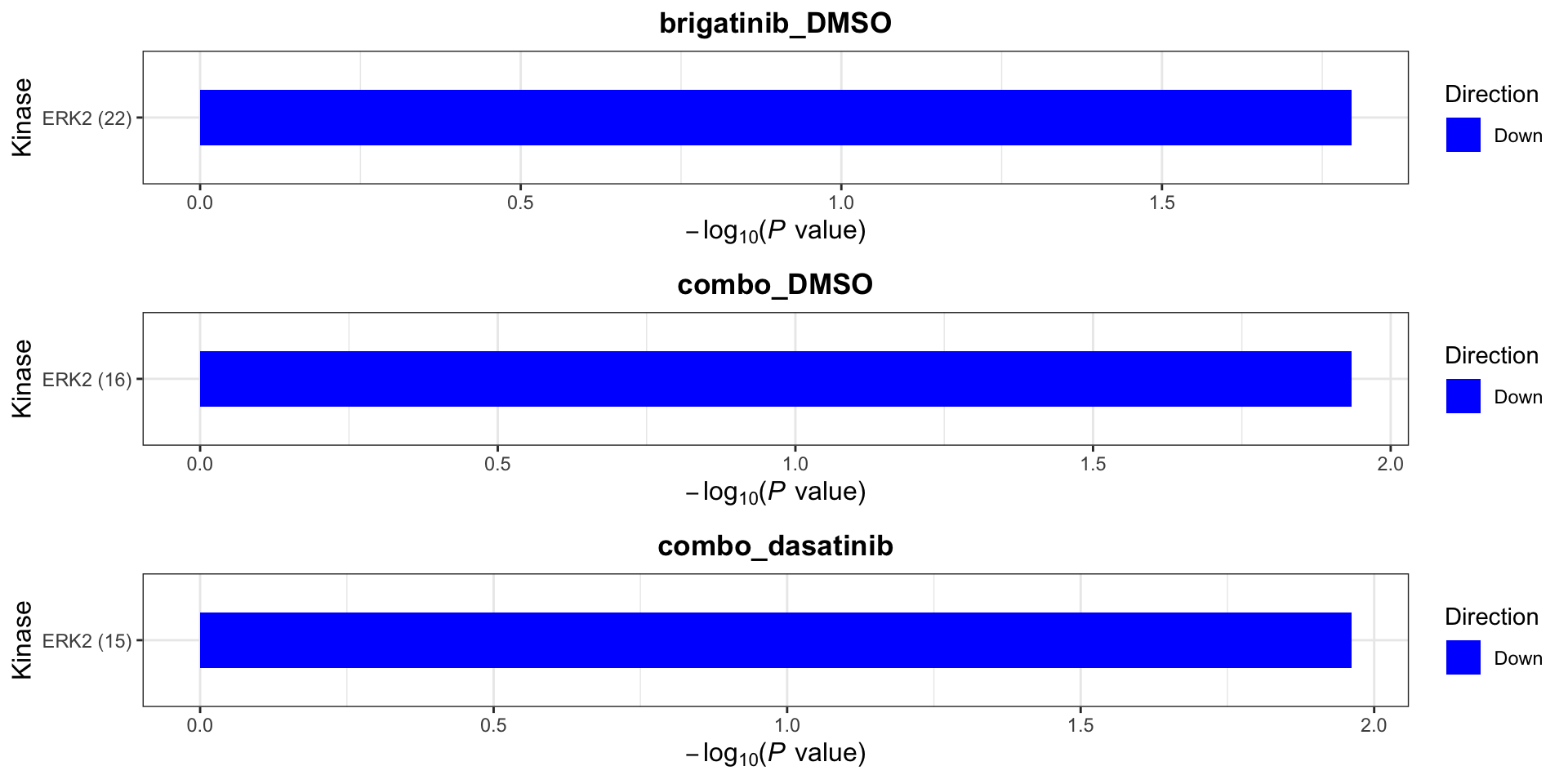
Site-specific Kinase enrichment
Here the site-specificity is considered.
P=0.01 as cut-off
resTab.site <- mutate(resTab, symbol = site)
plotList <- runGeneSetEnrichment(resTab.site, kins, genePCut = 0.05, pCutSet = 0.01, setFdr = FALSE, method="fisher")[1] "Testing for: Kinase"
[1] "Condition: dasatinib_DMSO"
[1] "Condition: brigatinib_DMSO"
[1] "Condition: combo_DMSO"
[1] "Condition: combo_brigatinib"
[1] "Condition: combo_dasatinib"plotList$Kinase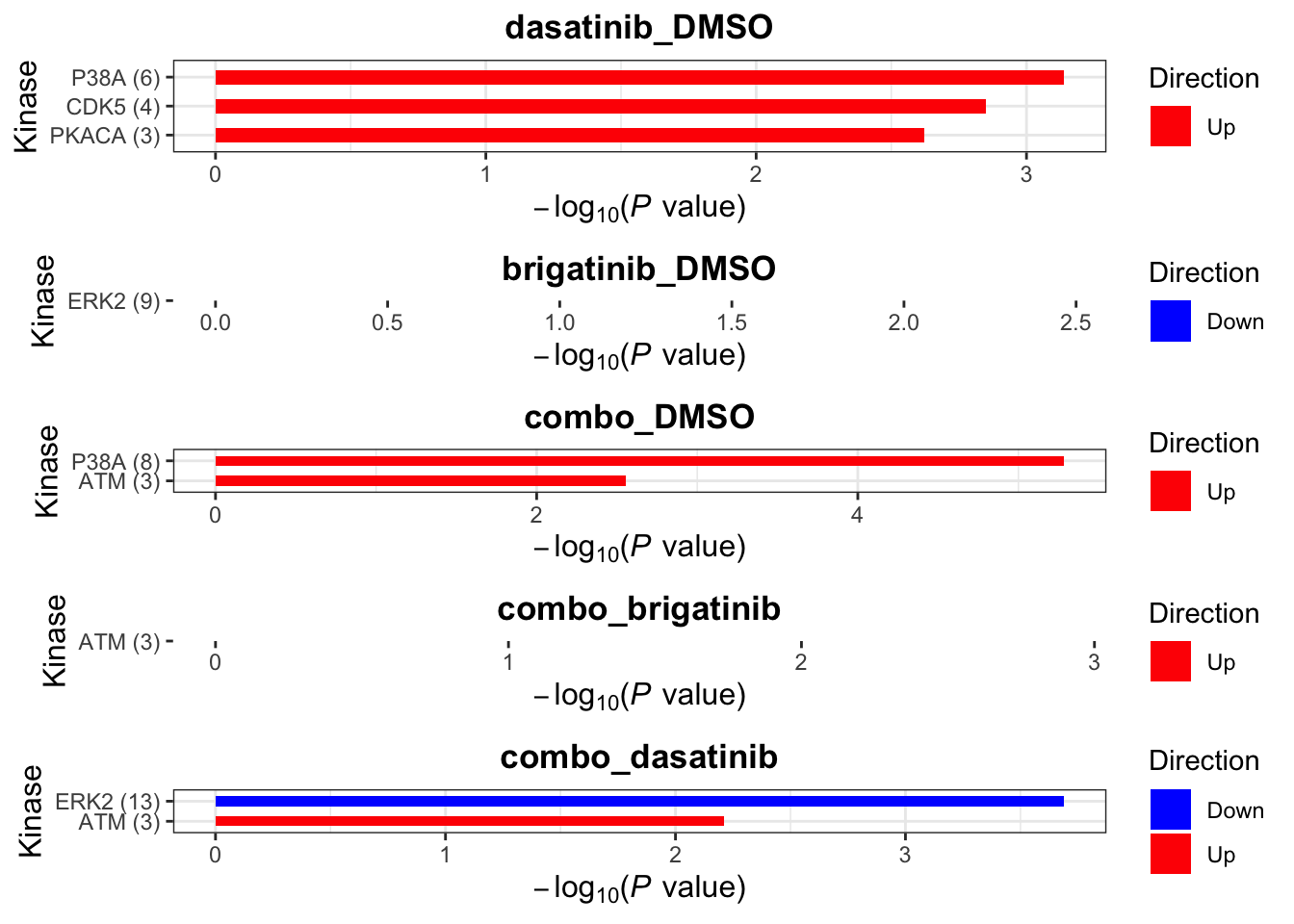
Focuse on the combination effect between Brigatinib and Dasatinib (Interaction effect)
Since Brigatinib and Dasatinib has synergistic effect on cell survival, this part is to detect the synergistic effect also on protein expression level. In statistical term, this is an interaction effect, where the effect of two variables is beyond symbol linear additive effect.
Differential test
Used save results, remove mitochondrial genes
resTab <- allResList$diffPhos$time_16 %>% filter(compare =="interaction")
resTab.sig <- filter(resTab, adj_pval <= 0.01)Here I used 0.01 adjusted p-values, otherwise there will be too many proteins
PCA plot with proteins showing interactions
PCA
PC1 versus PC2
plotPCA(fpeSub[resTab.sig$name,], assayName = "imputed", "PC1", "PC2", label ="replicate")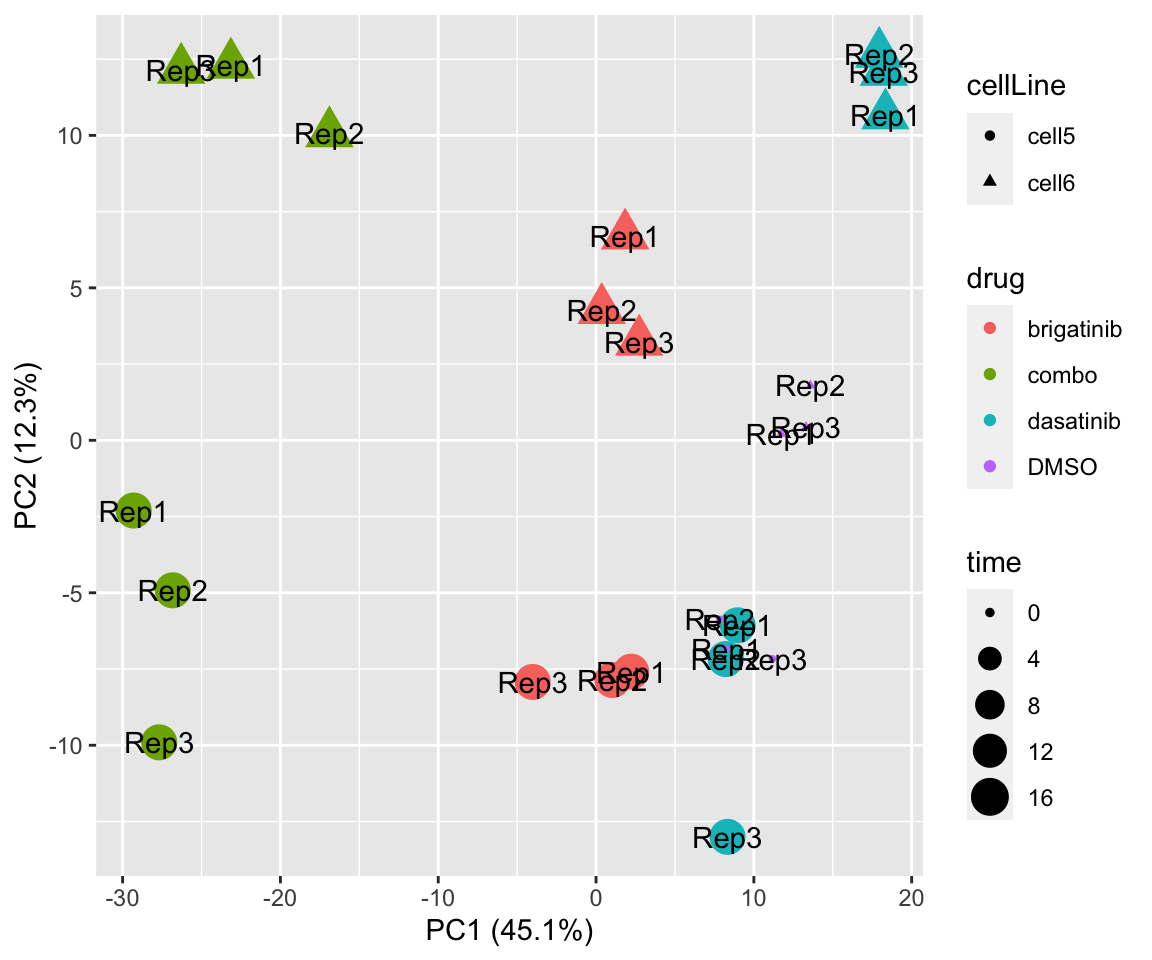 Potential problem with 1 replicate
Potential problem with 1 replicate
PC3 versus PC4
plotPCA(fpeSub[resTab.sig$name,], assayName = "imputed", "PC3", "PC4", label ="replicate")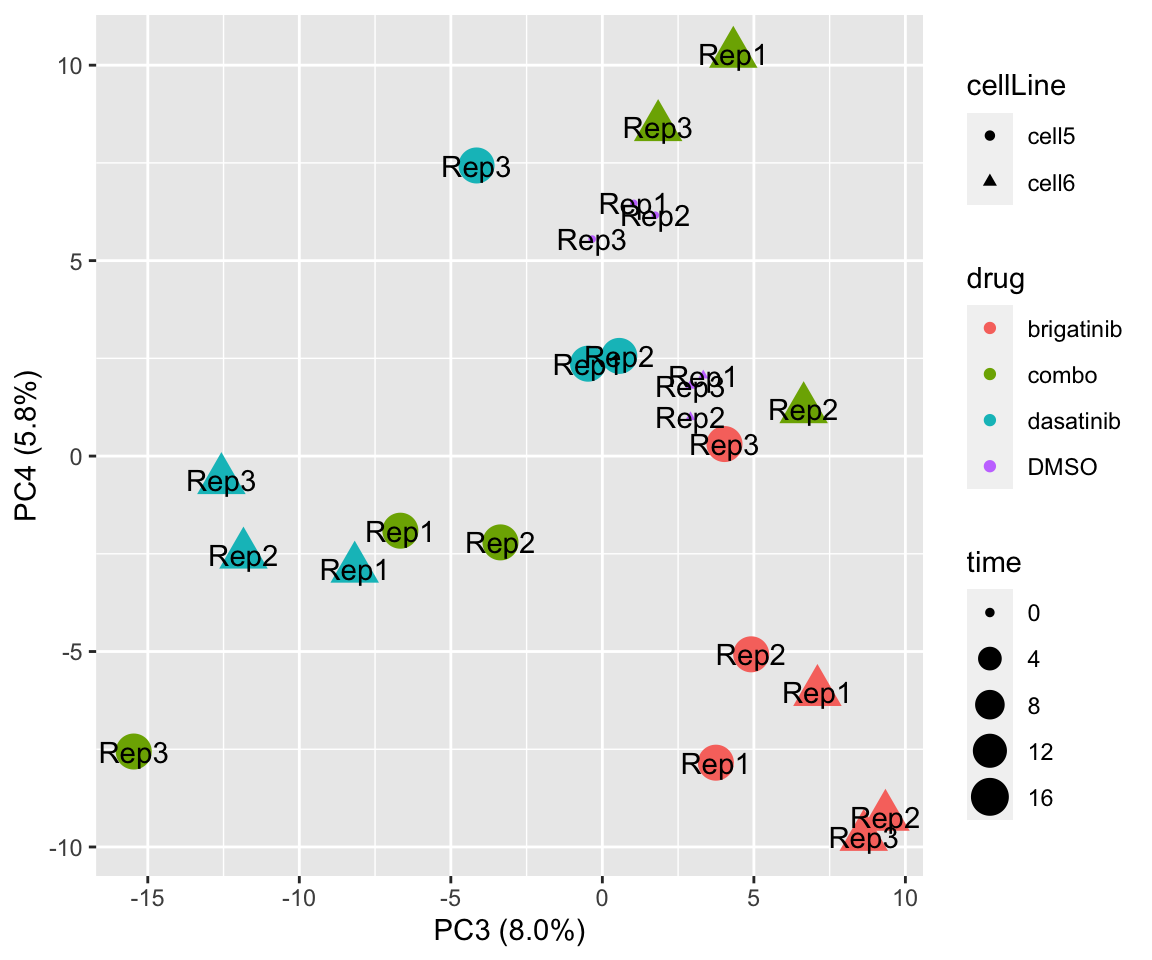
List of proteins with significant interactions (10%FDR)
resTab.sig %>% mutate(across(where(is.numeric), formatC, digits=2)) %>% DT::datatable()Heatmap of top 100 examples
plotProteinHeatmap(resTab, fpeSub, "all", fdrCut = 0.1, ifFDR = TRUE)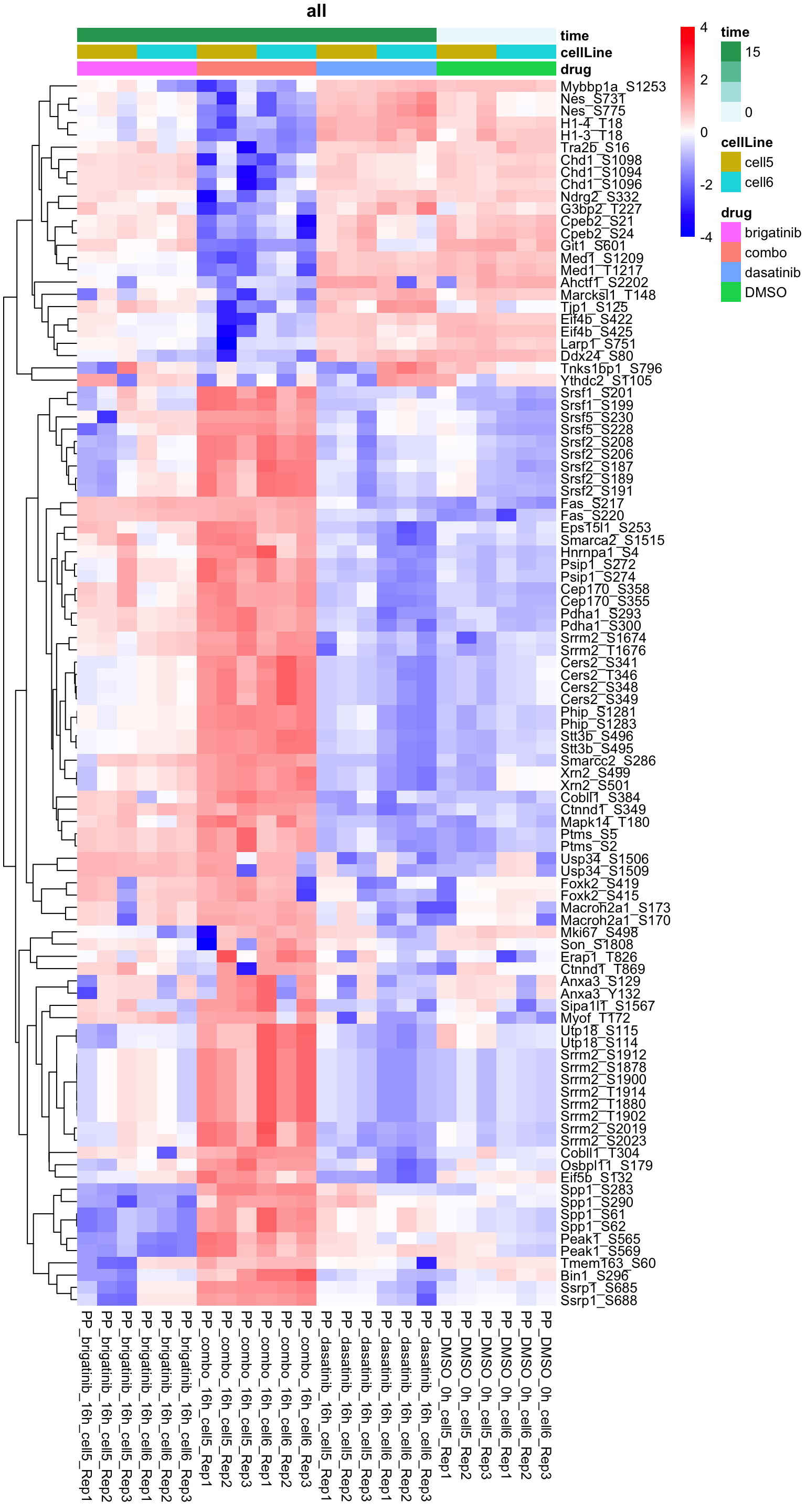
Box-Plot top 20 examples
plotTab <- jyluMisc::sumToTidy(fpeSub)
plotList <- lapply(seq(nrow(resTab.sig)), function(i) {
rec <- resTab.sig[i,]
eachTab <- filter(plotTab, rowID == rec$name)
ggplot(eachTab, aes(x=drug, y=Intensity)) +
geom_boxplot(aes(fill = drug)) +
ggbeeswarm::geom_quasirandom() +
facet_wrap(~cellLine) +
theme(legend.position = "none") +
ggtitle(rec$site)
})
cowplot::plot_grid(plotlist = plotList[1:20], ncol=2)
#plot all case in pdf file
#jyluMisc::makepdf(plotList, "../docs/boxplot_interactionPhos_16_RUN5.pdf", ncol = 2, nrow =5, height = 15, width = 10)Gene set enrichment analysis
Define useful genesets
plotList <- runGeneSetEnrichment(resTab, gmts, genePCut = 1, pCutSet = 0.05, geneFdr = FALSE, setFdr = TRUE, method="gsea", collapsePathway = TRUE)[1] "Testing for: Hallmark"
[1] "Condition: interaction"
[1] "No sets passed the criteria"
[1] "Testing for: CanonicalPathway"
[1] "Condition: interaction"
[1] "No sets passed the criteria"
[1] "Testing for: Kinase"
[1] "Condition: interaction"Hallmarks
plotList$Hallmark[1] "No sets passed the criteria"Canonical pathways
plotList$CanonicalPathway[1] "No sets passed the criteria"List of leading edge genes for each pathway
DT::datatable(plotList$leadingEdgeGene)writexl::write_xlsx(plotList$leadingEdgeGene, path = "../docs/enrichment_tables/phos_gsea_16combo_all.xlsx")Site-unspecific Kinase enrichment
On protein level, site specificity is not considered. This is because many phosphorylation sites lack the up-stream kinase annotation in the database. We can use a less stringent criteria to define kinase-substrate relation ship
plotList$Kinase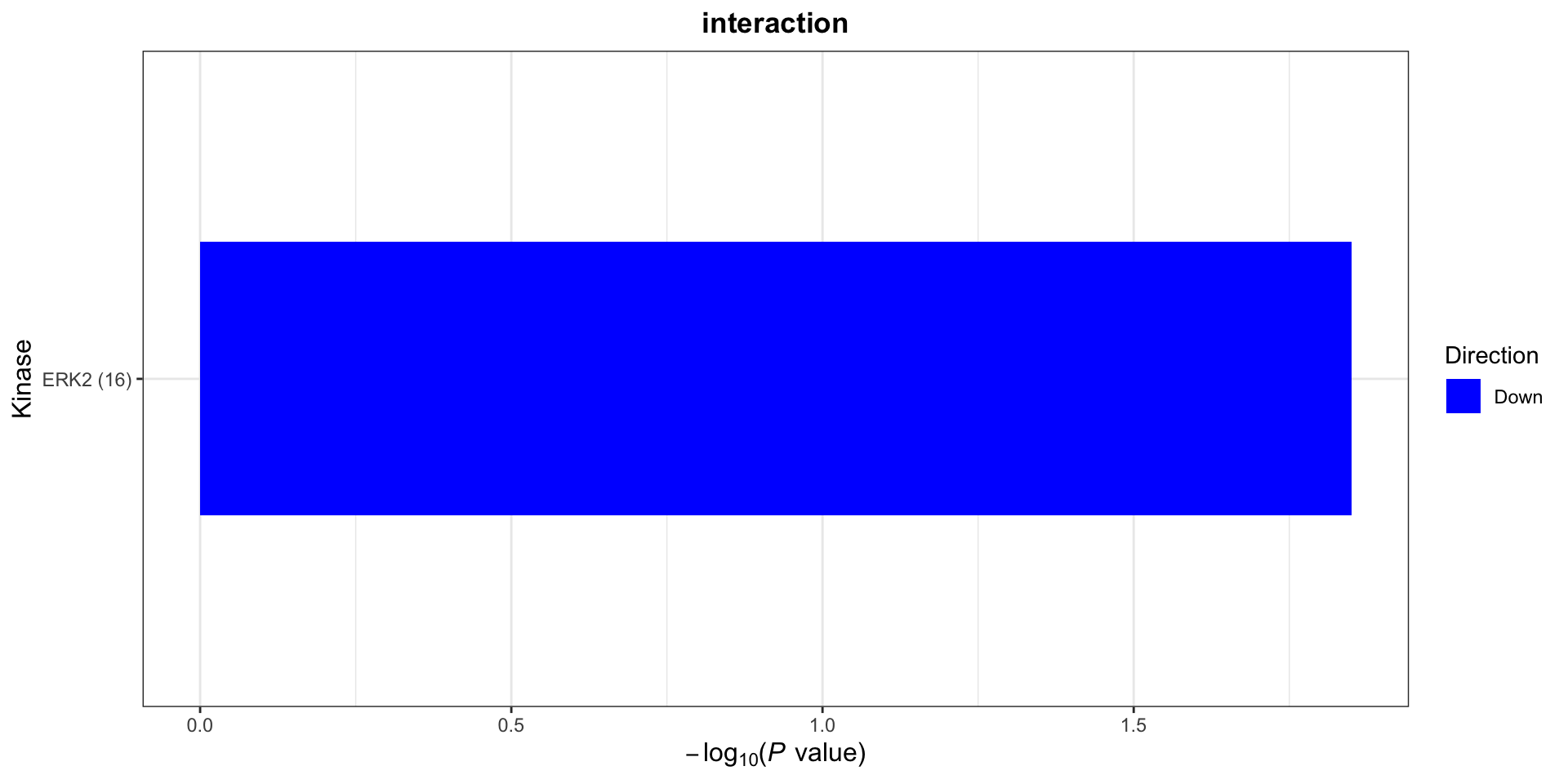
Site-specific Kinase enrichment
Here the site-specificity is considered.
P=0.01 as cut-off
resTab.site <- mutate(resTab, symbol = site)
plotList <- runGeneSetEnrichment(resTab.site, kins, genePCut = 0.05, pCutSet = 0.01, setFdr = FALSE, method="fisher")[1] "Testing for: Kinase"
[1] "Condition: interaction"plotList$Kinase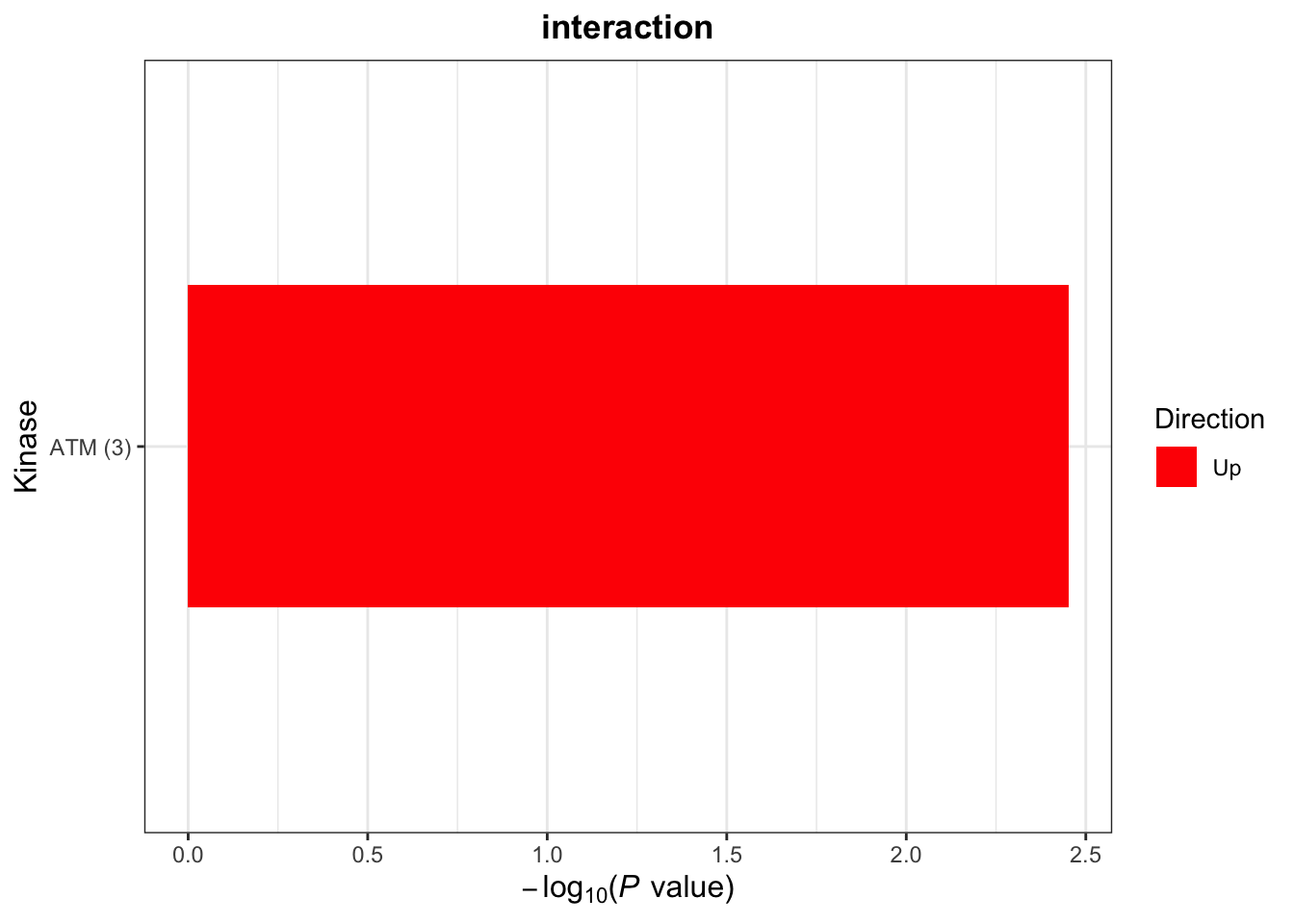
Plot interesting pathways
PI3K-AKT-MTOR
All conditions will be included, difference among cell lines will be regressed out for better visualization
plotSetHeatmap(fpeSub, filter(resTab, pval < 0.05), drugPair = "all", gmtFile = gmts$Hallmark,
setName = "HALLMARK_PI3K_AKT_MTOR_SIGNALING")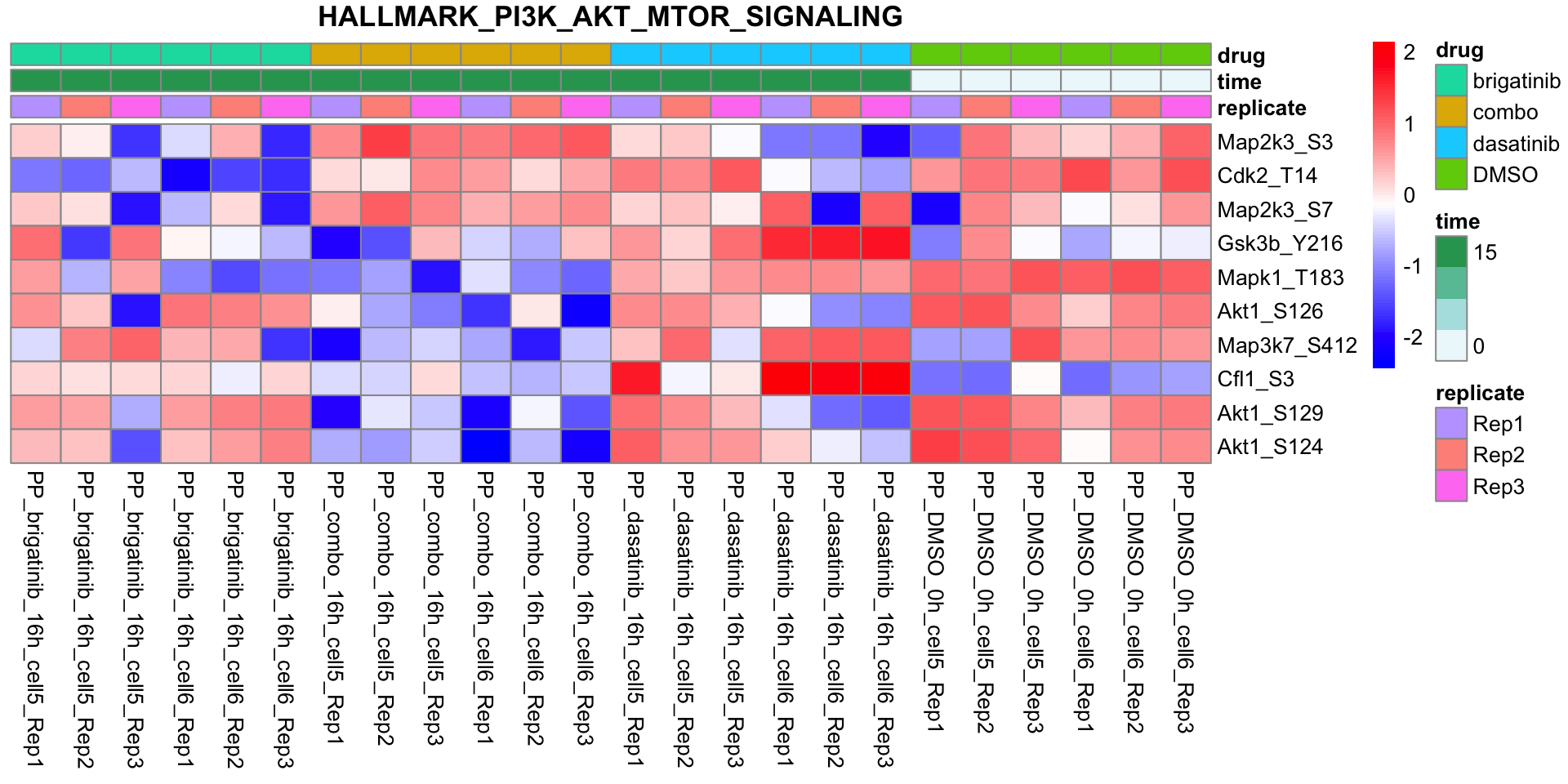
G2M_CHECKPOINT
All conditions will be included, difference among cell lines will be regressed out for better visualization
plotSetHeatmap(fpeSub, filter(resTab, pval < 0.05), drugPair = "all", gmtFile = gmts$Hallmark,
setName = "HALLMARK_G2M_CHECKPOINT")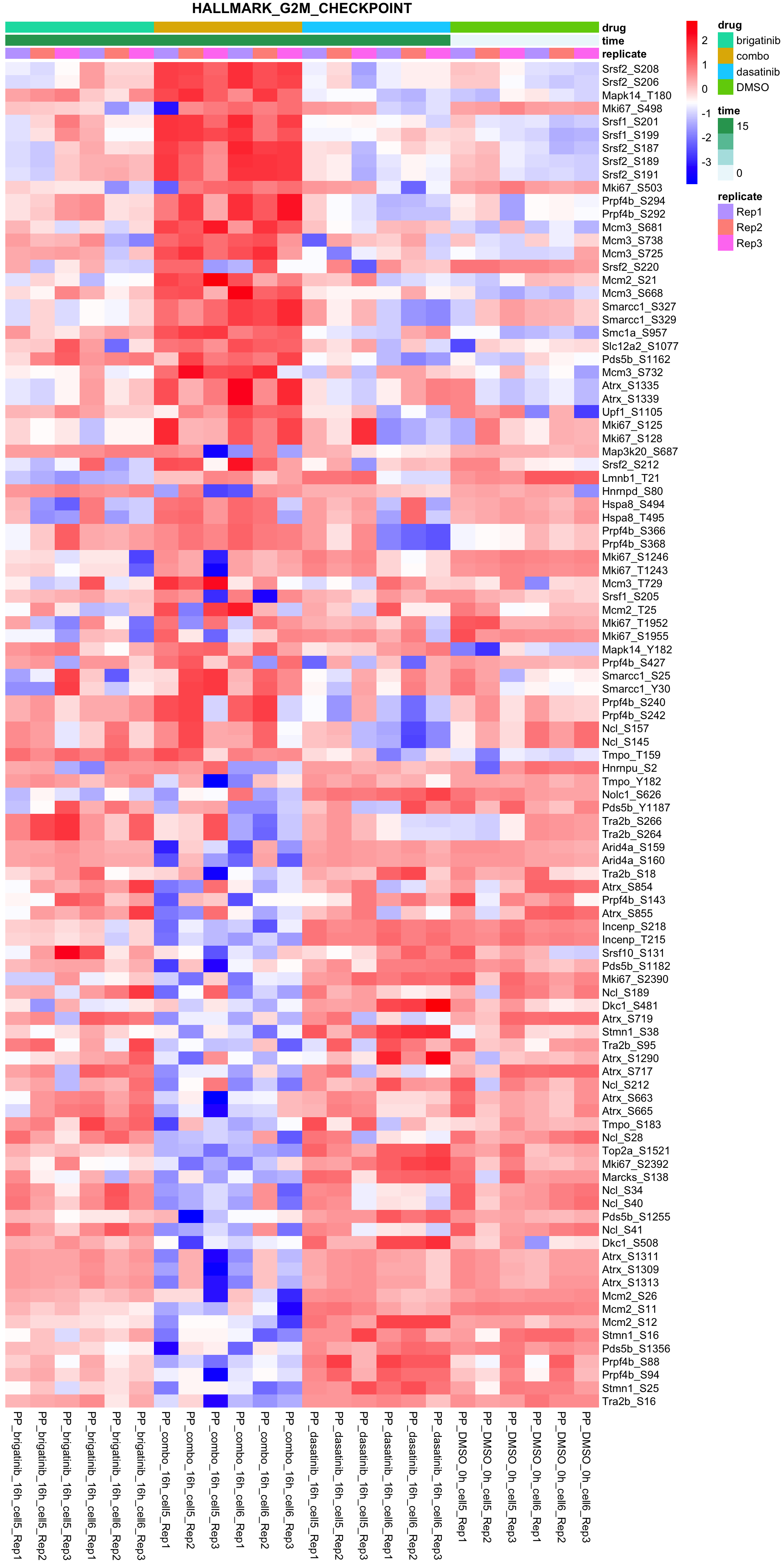
Create excel tables for download
Processed phosphorylation table
Normalized and log transformed. No imputation
seObj <- maeData[["Phosphoproteome"]]
protAnno <- rowData(seObj) %>% as_tibble(rownames="id") %>%
mutate(onMitochondria = ifelse(Gene %in% mitoList,"yes","no")) %>%
select(id,Gene, Residue, Position, onMitochondria) %>%
mutate(Site = paste0(Gene,"_",Residue,Position))
protMat <- assay(seObj)
protMat <- vsn::justvsn(protMat)
protTab <- protAnno %>% left_join(as_tibble(protMat, rownames = "id"), by = "id") %>%
dplyr::select(-id)
writexl::write_xlsx(protTab, path = "../docs/phosphorylation_matrix_RUN5.xlsx")Differential expression test results
output <- lapply(allResList$diffPhos,
function(x) {
x <- x %>% dplyr::rename(logFoldChange = diff) %>%
dplyr::select(-name)
x
})
writexl::write_xlsx(output, path = "../docs/dePhosRes_RUN5.xlsx")Heatmap plot of selected pathways
Prepare protein expression data
filterList <- list(time = c(0.17,0,16))
fpeSub <- preprocessPhos(maeData, filterList, missCut = 1, transform = "vst", normalize = TRUE)[1] "Number of proteins and samples:"
[1] 17182 42Ferroptosis
none of the genes passed the criteria
Metabolic, glycolysis, OXPHOS
Select genes to plot
seleGene <- read_csv2("../data/metabolic_glyco_oxphos.csv")
resTabList <- lapply(unique(seleGene$cluster2), function(x) {
geneList <- filter(seleGene, cluster2 == x)$gene
bind_rows(allResList$diffPhos$time_0.17, allResList$diffPhos$time_16) %>%
filter(compare %in% c("dasatinib_DMSO","brigatinib_DMSO", "combo_DMSO"),
pval <= 0.05,
symbol %in% geneList) %>%
distinct(name, .keep_all = TRUE)
})
names(resTabList) <- unique(seleGene$cluster2)Metabolic pathway
plotProteinListHeatmap(resTabList$Metabolic_pathway, fpeSub, title = "Metabolic_pathway")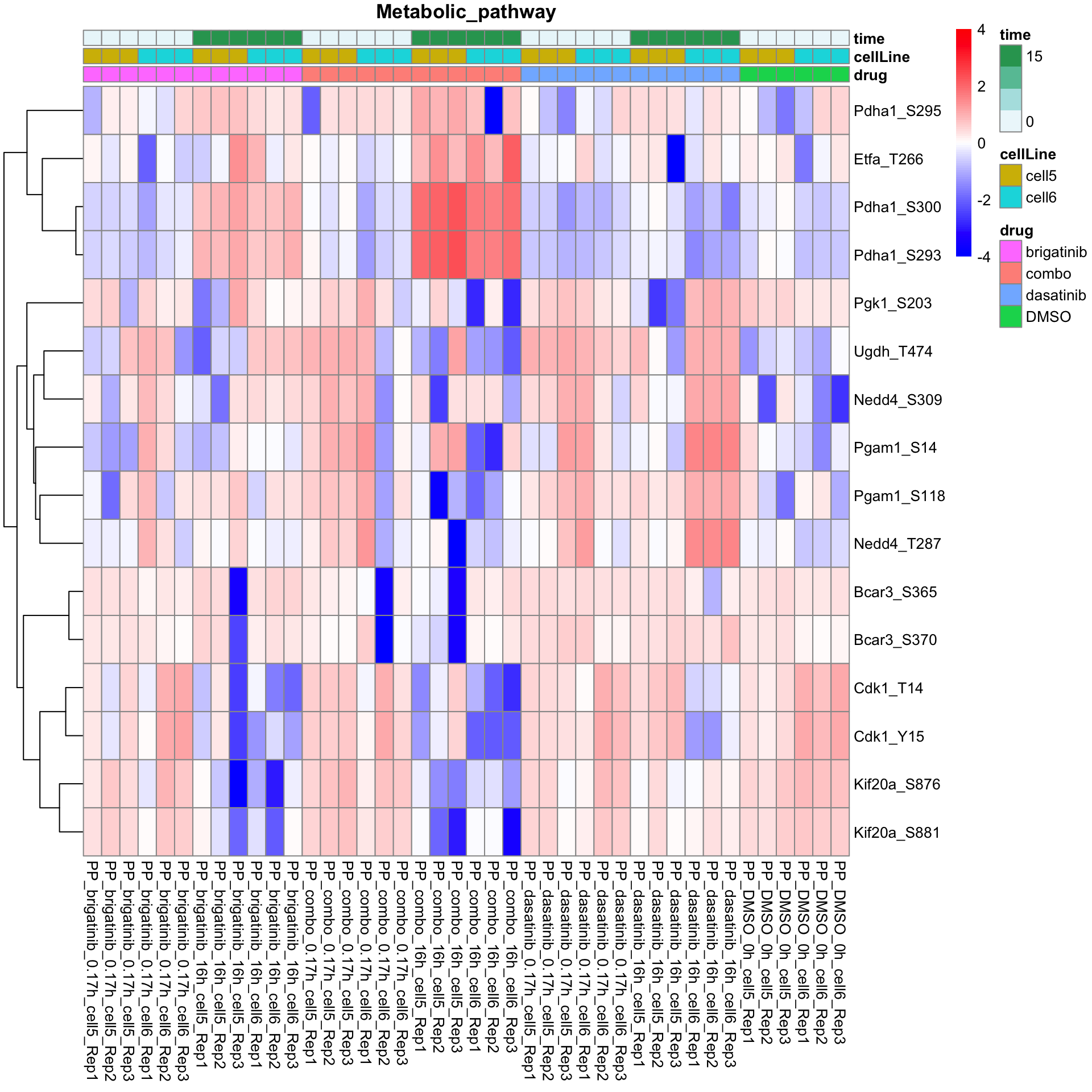
OXPHOS
plotProteinListHeatmap(resTabList$OXPHOS, fpeSub, title = "OXPHOS")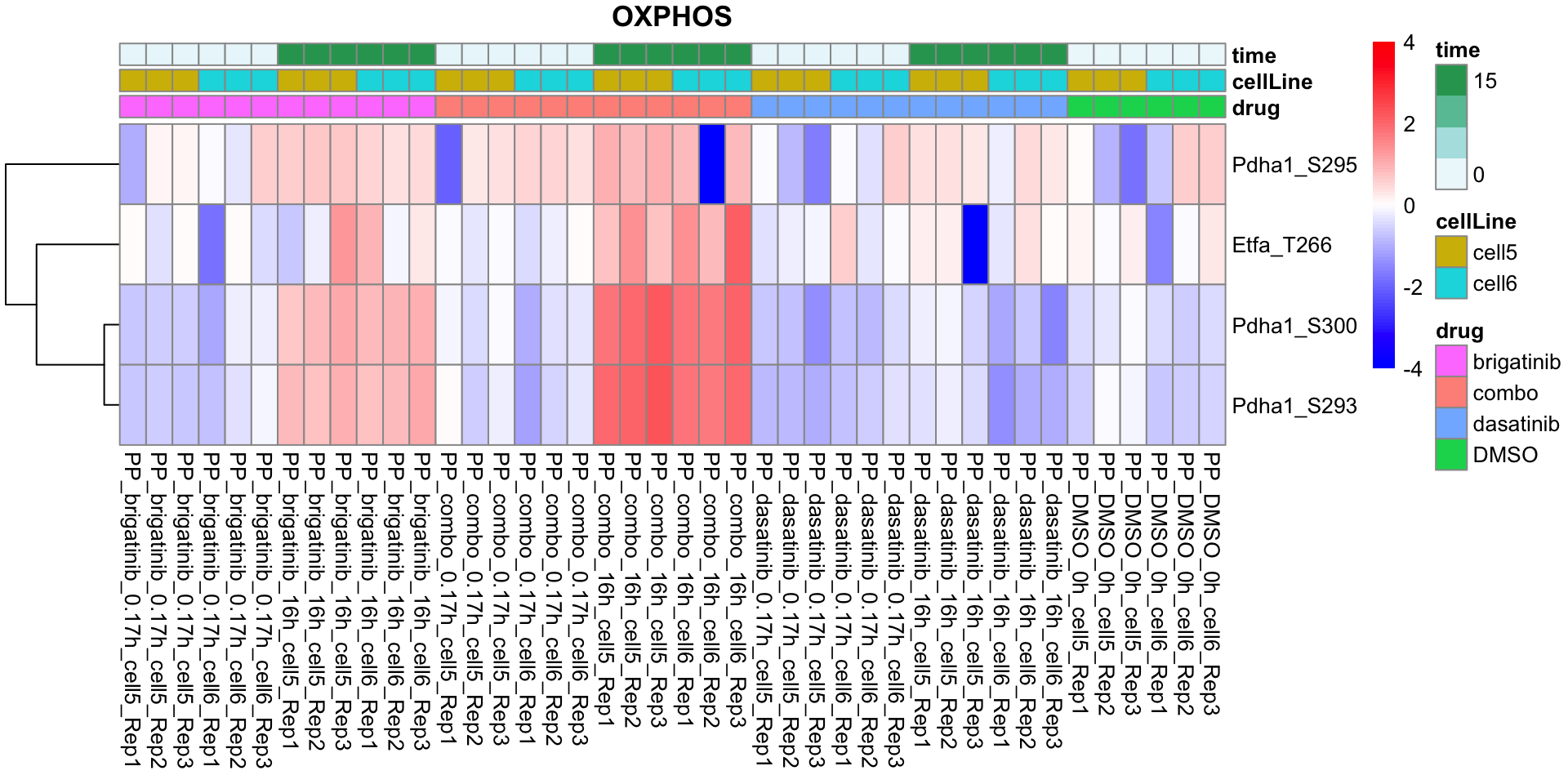
Glycolysis
plotProteinListHeatmap(resTabList$Glycolysis, fpeSub, title = "Glycolysis")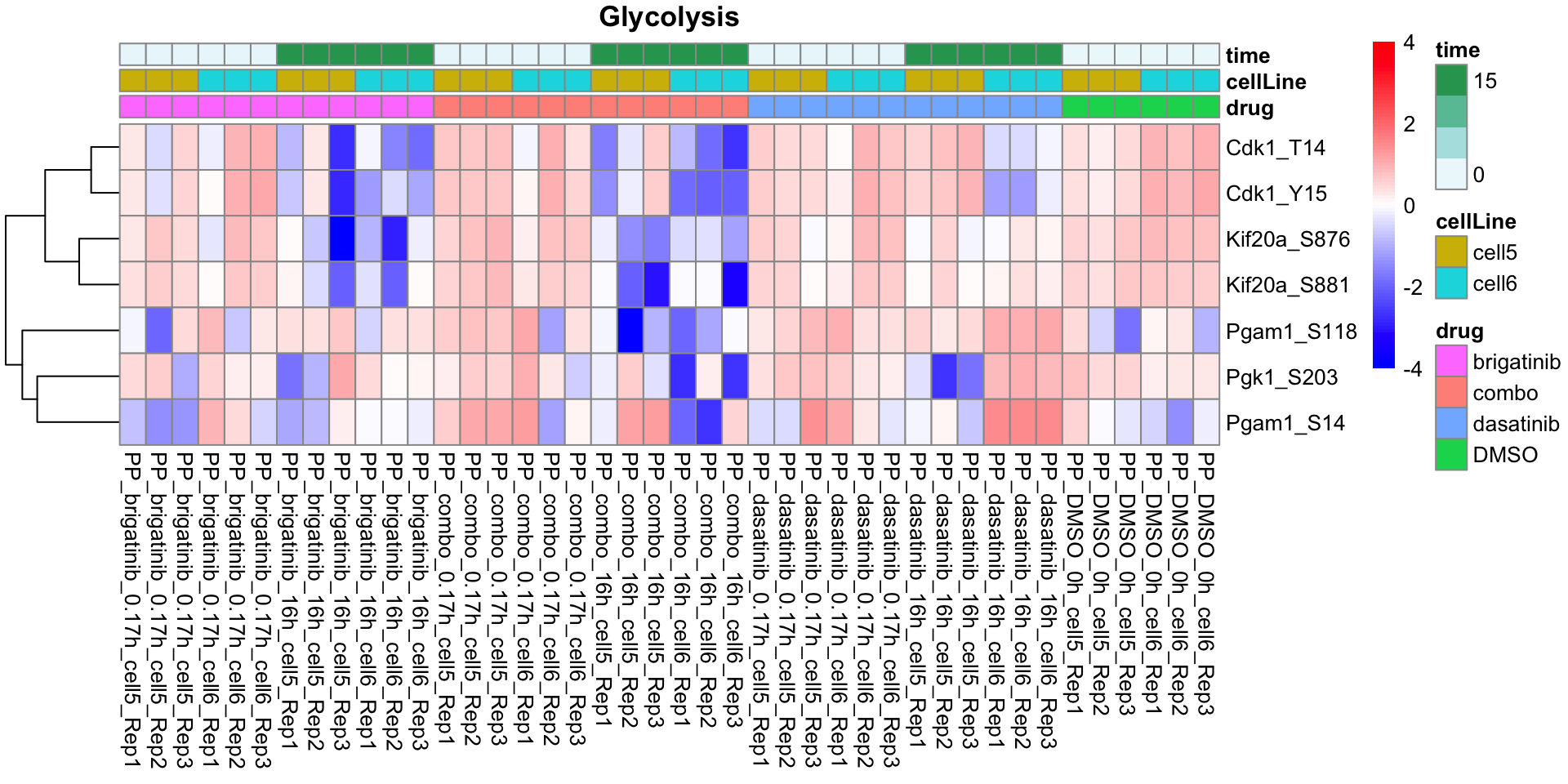
MYC pathway
Select genes to plot
seleGene <- bind_rows(tibble(gene = piano::loadGSC("../data/gmts/c2.cp.pid.v2022.1.Hs.symbols.gmt")$gsc$PID_MYC_ACTIV_PATHWAY, cluster2 = "MYC_ACTIVE_PATHWAY"),
tibble(gene = piano::loadGSC("../data/gmts/c2.cp.pid.v2022.1.Hs.symbols.gmt")$gsc$PID_MYC_REPRESS_PATHWAY, cluster2 = "PID_MYC_REPRESS_PATHWAY"))
resTabList <- lapply(unique(seleGene$cluster2), function(x) {
geneList <- filter(seleGene, cluster2 == x)$gene
bind_rows(allResList$diffPhos$time_0.17, allResList$diffPhos$time_16) %>%
filter(compare %in% c("dasatinib_DMSO","brigatinib_DMSO", "combo_DMSO"),
pval <= 0.05,
toupper(symbol) %in% geneList) %>%
distinct(name, .keep_all = TRUE)
})
names(resTabList) <- unique(seleGene$cluster2)MYC active pathway
plotProteinListHeatmap(resTabList$MYC_ACTIVE_PATHWAY, fpeSub, title = "MYC_ACTIVE_PATHWAY")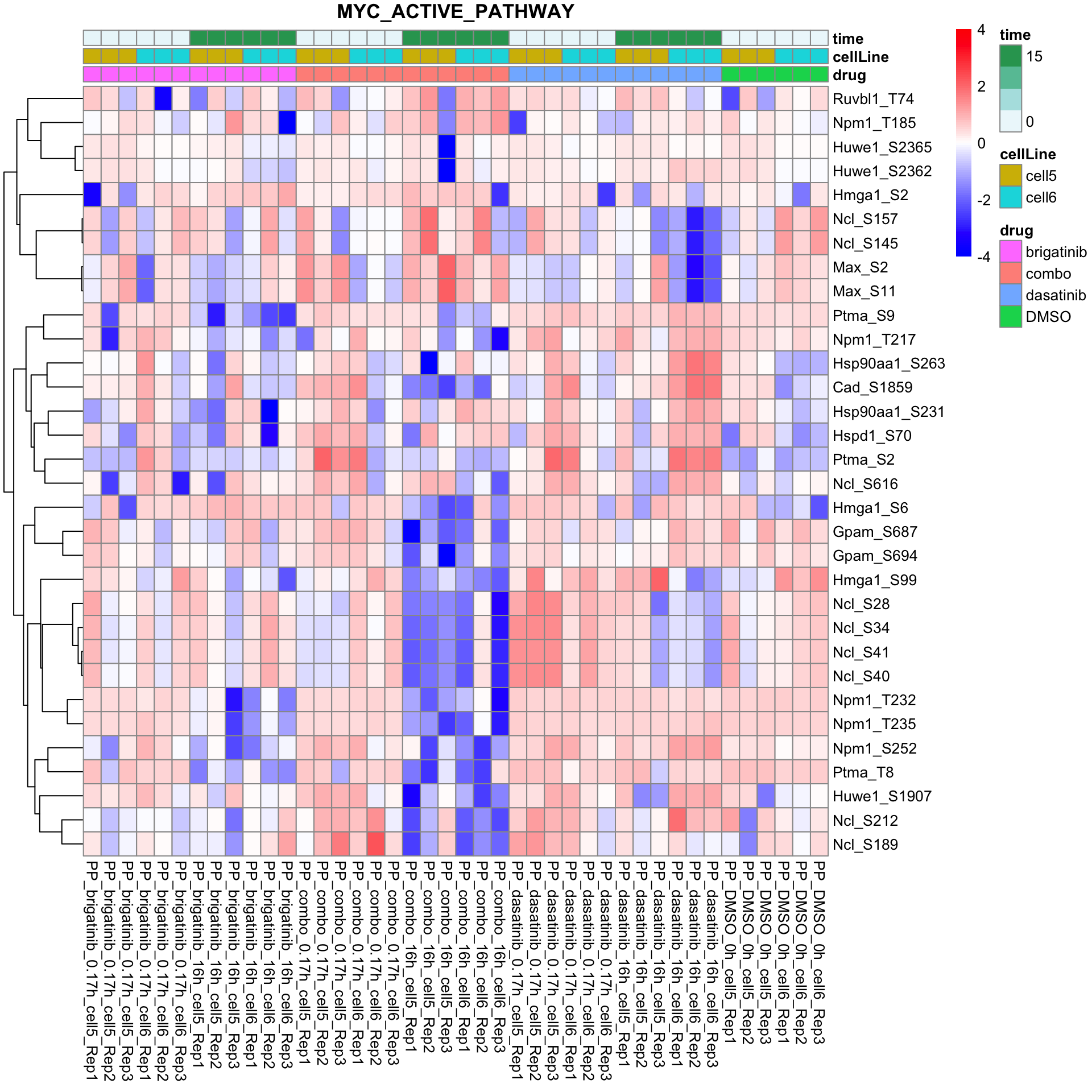
MYC repress pathway
plotProteinListHeatmap(resTabList$PID_MYC_REPRESS_PATHWAY, fpeSub, title = " MYC repress pathway")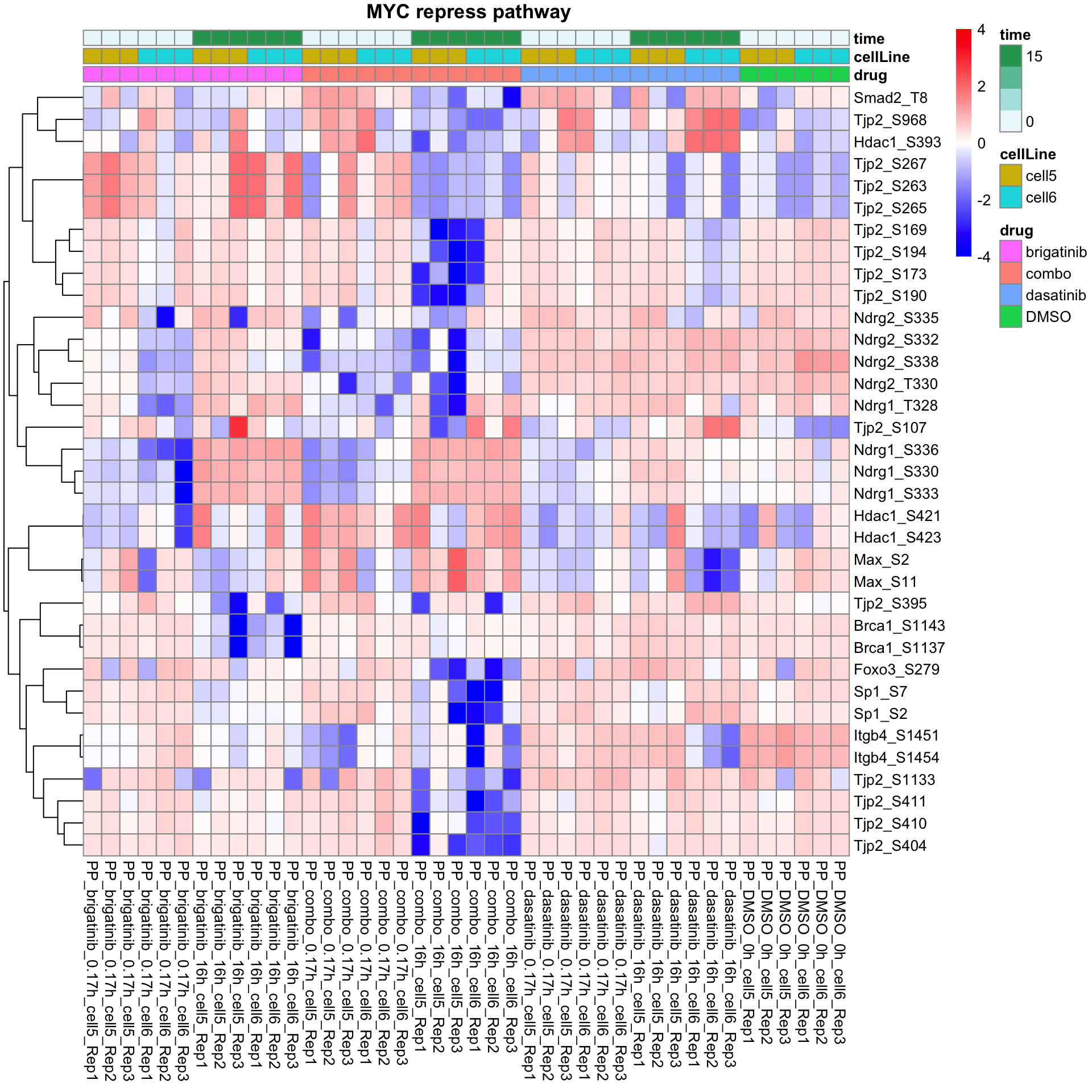
sessionInfo()R version 4.2.0 (2022-04-22)
Platform: x86_64-apple-darwin17.0 (64-bit)
Running under: macOS Big Sur/Monterey 10.16
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/4.2/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.2/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] stats4 stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] forcats_0.5.1 stringr_1.4.1
[3] dplyr_1.0.9 purrr_0.3.4
[5] readr_2.1.2 tidyr_1.2.0
[7] tibble_3.1.8 ggplot2_3.4.1
[9] tidyverse_1.3.2 UpSetR_1.4.0
[11] proDA_1.10.0 MultiAssayExperiment_1.22.0
[13] SummarizedExperiment_1.26.1 Biobase_2.56.0
[15] GenomicRanges_1.48.0 GenomeInfoDb_1.32.2
[17] IRanges_2.30.0 S4Vectors_0.34.0
[19] BiocGenerics_0.42.0 MatrixGenerics_1.8.1
[21] matrixStats_0.62.0
loaded via a namespace (and not attached):
[1] exactRankTests_0.8-35 bit64_4.0.5 knitr_1.39
[4] multcomp_1.4-19 DelayedArray_0.22.0 data.table_1.14.2
[7] KEGGREST_1.36.3 RCurl_1.98-1.7 doParallel_1.0.17
[10] generics_0.1.3 preprocessCore_1.58.0 cowplot_1.1.1
[13] TH.data_1.1-1 RSQLite_2.2.15 bit_4.0.4
[16] tzdb_0.3.0 xml2_1.3.3 lubridate_1.8.0
[19] httpuv_1.6.6 assertthat_0.2.1 gargle_1.2.0
[22] xfun_0.31 hms_1.1.1 jquerylib_0.1.4
[25] evaluate_0.15 promises_1.2.0.1 fansi_1.0.3
[28] caTools_1.18.2 dbplyr_2.2.1 readxl_1.4.0
[31] km.ci_0.5-6 igraph_1.3.4 DBI_1.1.3
[34] htmlwidgets_1.5.4 googledrive_2.0.0 ellipsis_0.3.2
[37] jyluMisc_0.1.5 crosstalk_1.2.0 ggpubr_0.4.0
[40] backports_1.4.1 annotate_1.74.0 vctrs_0.5.2
[43] imputeLCMD_2.1 abind_1.4-5 cachem_1.0.6
[46] withr_2.5.0 vroom_1.5.7 cluster_2.1.3
[49] crayon_1.5.2 drc_3.0-1 relations_0.6-12
[52] genefilter_1.78.0 edgeR_3.38.1 pkgconfig_2.0.3
[55] slam_0.1-50 labeling_0.4.2 nlme_3.1-158
[58] vipor_0.4.5 ProtGenerics_1.28.0 rlang_1.0.6
[61] lifecycle_1.0.3 sandwich_3.0-2 affyio_1.66.0
[64] modelr_0.1.8 cellranger_1.1.0 rprojroot_2.0.3
[67] Matrix_1.4-1 KMsurv_0.1-5 carData_3.0-5
[70] zoo_1.8-10 DEP_1.18.0 reprex_2.0.1
[73] beeswarm_0.4.0 GlobalOptions_0.1.2 googlesheets4_1.0.0
[76] pheatmap_1.0.12 png_0.1-7 rjson_0.2.21
[79] mzR_2.30.0 bitops_1.0-7 shinydashboard_0.7.2
[82] KernSmooth_2.23-20 visNetwork_2.1.0 Biostrings_2.64.0
[85] blob_1.2.3 workflowr_1.7.0 shape_1.4.6
[88] maxstat_0.7-25 rstatix_0.7.0 tmvtnorm_1.5
[91] ggsignif_0.6.3 scales_1.2.0 memoise_2.0.1
[94] magrittr_2.0.3 plyr_1.8.7 gplots_3.1.3
[97] zlibbioc_1.42.0 compiler_4.2.0 RColorBrewer_1.1-3
[100] plotrix_3.8-2 pcaMethods_1.88.0 clue_0.3-61
[103] cli_3.4.1 affy_1.74.0 XVector_0.36.0
[106] MASS_7.3-58 mgcv_1.8-40 tidyselect_1.1.2
[109] vsn_3.64.0 stringi_1.7.8 highr_0.9
[112] yaml_2.3.5 locfit_1.5-9.6 norm_1.0-10.0
[115] MALDIquant_1.21 survMisc_0.5.6 grid_4.2.0
[118] sass_0.4.2 fastmatch_1.1-3 tools_4.2.0
[121] parallel_4.2.0 circlize_0.4.15 rstudioapi_0.13
[124] MsCoreUtils_1.8.0 foreach_1.5.2 git2r_0.30.1
[127] gridExtra_2.3 farver_2.1.1 mzID_1.34.0
[130] digest_0.6.30 BiocManager_1.30.18 shiny_1.7.4
[133] Rcpp_1.0.9 car_3.1-0 broom_1.0.0
[136] later_1.3.0 writexl_1.4.0 ncdf4_1.19
[139] httr_1.4.3 survminer_0.4.9 MSnbase_2.22.0
[142] AnnotationDbi_1.58.0 ComplexHeatmap_2.12.0 colorspace_2.0-3
[145] rvest_1.0.2 XML_3.99-0.10 fs_1.5.2
[148] splines_4.2.0 gmm_1.6-6 xtable_1.8-4
[151] jsonlite_1.8.3 marray_1.74.0 R6_2.5.1
[154] sets_1.0-21 pillar_1.8.0 htmltools_0.5.4
[157] mime_0.12 glue_1.6.2 fastmap_1.1.0
[160] DT_0.23 BiocParallel_1.30.3 codetools_0.2-18
[163] fgsea_1.22.0 mvtnorm_1.1-3 utf8_1.2.2
[166] lattice_0.20-45 bslib_0.4.1 sva_3.44.0
[169] ggbeeswarm_0.6.0 gtools_3.9.3 shinyjs_2.1.0
[172] survival_3.4-0 limma_3.52.2 rmarkdown_2.14
[175] munsell_0.5.0 GetoptLong_1.0.5 GenomeInfoDbData_1.2.8
[178] iterators_1.0.14 piano_2.12.0 impute_1.70.0
[181] haven_2.5.0 gtable_0.3.0